-
Computational Biomedicine (CBM, Online ISSN: 3107-3131) is a peer-reviewed, open access journal published quarterly and owned by Science Exploration Press. The journal covers a wide range of topics, including molecular medicine, simulation, modeling techniques, imaging methods, and information technology. Our mission is to encourage scientists to publish their experimental and theoretical findings in a detailed open-access format. We invite submissions across various article types, including Research Articles, Review Articles, Editorials, Case Reports, Letters to the Editor, Perspectives, and Commentaries. more >
Articles
The application of attention mechanisms in biological sequence analysis
-
In recent years, attention mechanisms have gained widespread application and significant advancements in the field of biological sequence analysis. This paper systematically summarizes the fundamental principles of attention mechanisms and their latest ...
MoreIn recent years, attention mechanisms have gained widespread application and significant advancements in the field of biological sequence analysis. This paper systematically summarizes the fundamental principles of attention mechanisms and their latest research progress in biological sequence analysis. First, the development history of attention mechanisms is introduced, with a focus on classic mechanisms such as self-attention, cross-attention, and multi-head attention, along with their improved variants. Next, a brief overview of the classification and characteristics of bioinformatics databases is provided. Subsequently, the application of attention mechanisms in the analysis of DNA, RNA, and protein sequences is highlighted. In the realm of DNA sequence analysis, attention mechanisms have been applied to tasks such as epigenetic analysis and regulatory element identification; in RNA sequence analysis, they play a crucial role in single-cell RNA sequencing, RNA function prediction, and structure prediction; in protein sequence analysis, attention mechanisms are widely used in protein classification, function prediction, structure prediction, site prediction, and interaction prediction. Furthermore, this paper summarizes the applications of attention mechanisms in other biological sequence analysis tasks, such as multi-omics analysis and enzyme analysis. The attention mechanisms can significantly improve the accuracy, interpretability, and computational efficiency of biological sequence analysis, providing powerful computational tools for bioinformatics research.
Less -
Yingyue Tang, Wenzheng Bao
-
DOI: https://doi.org/10.70401/cbm.2026.0014 - May 15, 2026
PIONEER: A structure-informed graph neural network for PE/PPE protein identification
-
Aims: The Pro-Glu (PE) and Pro-Pro-Glu (PPE) protein family of Mycobacterium tuberculosis plays a critical role in virulence, immune evasion, and host-pathogen interactions. However, the high guanine-cytosine-content and repetitive ...
MoreAims: The Pro-Glu (PE) and Pro-Pro-Glu (PPE) protein family of Mycobacterium tuberculosis plays a critical role in virulence, immune evasion, and host-pathogen interactions. However, the high guanine-cytosine-content and repetitive sequences of these proteins have long hindered accurate gene identification and functional annotation. This study aims to develop an effective computational framework to improve the identification of PE/PPE proteins.
Methods: We propose PIONEER, a structure-aware deep learning framework that integrates embeddings from the pre-trained protein language model ESM Cambrian (ESMC) with structural features. PIONEER represents proteins as residue-level graphs that encode both sequence semantics and three-dimensional topological structure, enabling effective modelling of hierarchical geometric relationships.
Results: Benchmarking demonstrates that PIONEER outperforms 16 traditional machine learning algorithms and the existing deep learning model across multiple evaluation metrics, including accuracy, Matthew’s correlation coefficient, and F1 scores. Ablation experiments confirm the complementary contributions of ESMC embeddings and secondary structure features to model performance. The t-SNE-based visualization results reveal the contributions of features across different network layers to the identification of PE/PPE proteins.
Conclusion: PIONEER improves the accuracy of PE/PPE protein identification by integrating sequence and structural information within a structure-aware graph learning framework. This method provides an effective computational tool for functional annotation, investigation of pathogenicity mechanisms, and vaccine target discovery in M. tb.
Less -
Heyun Sun, ... Fuyi Li
-
DOI: https://doi.org/10.70401/cbm.2026.0016 - May 11, 2026
Distilling genomic knowledge into pathology slides for robust cancer survival prediction
-
Aims: To develop a robust and clinically feasible framework for cancer survival prediction using only histopathology images while leveraging transcriptomic knowledge during training.
Methods: The study proposed Adaptive Multi-modality ...
MoreAims: To develop a robust and clinically feasible framework for cancer survival prediction using only histopathology images while leveraging transcriptomic knowledge during training.
Methods: The study proposed Adaptive Multi-modality Knowledge Distillation (AMKD), a framework designed to transfer complementary molecular-level information from transcriptomic data to pathology-based models. The AMKD framework consists of two essential elements. First, a gene-guided pathology enhancement module is designed to inject genomics-aware information from a multimodal teacher into pathology features. Second, an adaptive redundancy reduction loss is introduced to regulate knowledge distillation by accounting for prediction discrepancies between teacher and student models. This design allows the student model to retain biologically meaningful knowledge during training and remain effective with only histopathology data at inference.
Results: Comprehensive experiments on four The Cancer Genome Atlas (TCGA) cancer cohorts demonstrate that AMKD achieves
state-of-the-art survival prediction performance, with an average concordance index (C-index) of 0.669, surpassing both unimodal pathology-based and fully multi-modal approaches. External validation on the independent Clinical Proteomic Tumor Analysis Consortium head and neck squamous cell carcinoma (CPTAC-HNSC) cohort further confirms the robustness and cross-dataset generalization ability of the proposed framework. Ablation studies further confirm the effectiveness of each proposed component in enhancing cross-modal knowledge transfer and improving generalization under incomplete modality scenarios.Conclusion: The proposed AMKD framework provides a clinically practical solution for robust cancer survival analysis when transcriptomic data are unavailable. By adaptively distilling multi-modal knowledge into a pathology-based model, AMKD bridges the gap between research and clinical applicability, enabling scalable and cost-effective prognostic prediction in real-world settings.
Less -
Yangfan Xu, ... Runming Wang
-
DOI: https://doi.org/10.70401/cbm.2026.0015 - April 29, 2026
A deep learning framework with positional attention for modeling enhancer-promoter interactions
-
Aims: Since distal enhancers are involved in regulating target genes through physical contacting with proximal promoters, identifying enhancer-promoter interactions (EPIs) is critical to deepening our understanding of gene expression. However, ...
MoreAims: Since distal enhancers are involved in regulating target genes through physical contacting with proximal promoters, identifying enhancer-promoter interactions (EPIs) is critical to deepening our understanding of gene expression. However, high-throughput experimental methods for identifying EPIs are time-consuming and expensive. Therefore, computational methods for predicting EPIs would be valuable and important, but also face a lot of challenges.
Methods: In this paper, we propose a novel deep learning-based method, namely EPIPAM, to predict EPIs only using genomic sequences. EPIPAM firstly uses a deep convolutional neural network to extract high-level sequence features, and then uses a position attention mechanism to compute the positional correlation coefficients of two subregions separately coming from enhancers and promoters, aiming to focus on important regions of them.
Results: Benchmarking comparisons on six different cell lines show that EPIPAM performs better than the state-of-the-art methods in the task of EPIs prediction. More importantly, we notice that, almost without exception, the predictive performance of all methods is really poor once applying a strategy of splitting training and test data by chromosome. Therefore, we explain the possible reason that leads to this situation by systematically exploring the structure of EPI datasets, and indirectly analyze the difficulty of predicting EPIs only using genomic sequences through ChIA-PET contact datasets.
Conclusion: This study presents a novel deep learning-based method to predict EPIs only using genomic sequences. Although the proposed method achieves higher predictive accuracy, it suffers from several limitations, such as highly selective matching bias, negative sample selection issues, and constraints of pre-trained vectors.
Less -
Liping Liu, ... Qinhu Zhang
-
DOI: https://doi.org/10.70401/cbm.2026.0013 - April 08, 2026
Multi-class pattern discovery for bacterial secretory effectors
-
Aims: EffecTri aims to develop a comprehensive, multi-class prediction framework to accurately identify bacterial effector proteins secreted by Type III, IV, and VI secretion systems. Current methodologies often employ binary classifications, ...
MoreAims: EffecTri aims to develop a comprehensive, multi-class prediction framework to accurately identify bacterial effector proteins secreted by Type III, IV, and VI secretion systems. Current methodologies often employ binary classifications, overlooking the complexity and interactions among multiple effector classes.
Methods: EffecTri integrates deep contextual embeddings from Evolutionary Scale Modeling and handcrafted descriptors, including Amino Acid Composition and Dipeptide Composition. The performance of the model was rigorously evaluated through comparative descriptor analyses and optimized feature combinations, complemented by Uniform Manifold Approximation and Projection visualization for interpretability.
Results: EffecTri outperformed traditional machine learning methods, achieving a weighted F1-score of 0.850 on an independent test dataset. The fusion of Evolutionary Scale Modeling embeddings with handcrafted descriptors demonstrated superior predictive performance, clearly distinguishing effector classes in UMAP visualizations.
Conclusion: EffecTri represents a robust, interpretable, and accurate computational tool, enhancing the multi-class identification of bacterial secretory effectors and contributing valuable insights into bacterial pathogenic mechanisms.
Less -
Jing Li, ... Youyu Wang
-
DOI: https://doi.org/10.70401/cbm.2026.0012 - March 05, 2026
A comprehensive review on neuropeptides: databases and computational tools
-
Neuropeptides are crucial signaling molecules that regulate diverse physiological processes spanning growth, social behavior, learning, memory, metabolism, homeostasis, reproduction, and neural differentiation across both nervous and peripheral ...
MoreNeuropeptides are crucial signaling molecules that regulate diverse physiological processes spanning growth, social behavior, learning, memory, metabolism, homeostasis, reproduction, and neural differentiation across both nervous and peripheral systems. Dysregulation of neuropeptides signaling is closely linked to various pathological conditions, such as neurological disorders, metabolic diseases, cardiovascular conditions, and even cancer, positioning them as potential therapeutic agents or targets for intervention. In recent years, research into neuropeptides has accelerated, with vast amounts of data continuously accumulating in multiple databases. However, the study of neuropeptides is often impeded by the need for extensive and time-consuming experimental investigations. As a result, computational tools have become essential for the rapid, large-scale identification of neuropeptides. This review systematically discusses neuropeptide-related databases and computational tools. These databases organize extensive data on neuropeptide sequences, structures, and functions. Among these, NeuroPep2.0, with 11,417 neuropeptide entries, is currently the most widely used dataset for neuropeptide prediction. Additionally, this review explores the application of computational approaches in neuropeptide prediction. While early methods predominantly relied on homologous sequence alignment and biochemical feature statistics, recent advances in machine learning have significantly enhanced prediction accuracy and efficiency. Tools such as NeuroPred-PLM and DeepNeuropePred, developed by our research group using protein language models, have substantially improved prediction performance. In conclusion, this review provides a comprehensive overview of current neuropeptide databases and computational tools, offering researchers a thorough survey of available resources and analytical methods, and emphasizing the necessity of continuous optimization to advance neuropeptide research and its therapeutic applications.
Less -
Wei Xu, ... Yan Wang
-
DOI: https://doi.org/10.70401/cbm.2025.0001 - April 10, 2025
MediHerb: A multi-modal enhanced framework for disease inference via herbal knowledge
-
Aims: Development of robust and effective methods for uncovering herb interactions and constructing herb–disease associations requires the integration of diverse biological and medical information. A key challenge in Traditional Chinese Medicine ...
MoreAims: Development of robust and effective methods for uncovering herb interactions and constructing herb–disease associations requires the integration of diverse biological and medical information. A key challenge in Traditional Chinese Medicine (TCM) research is to robustly uncover herb interactions and construct reliable herb–disease associations. This task requires handling the inherently high-dimensional, multi-label, and cross-domain nature of prescription data. Existing approaches provide limited representation capacity and insufficient integration of biomedical knowledge, restricting their ability to capture the complex semantics underlying the relationships between herbs and diseases.
Methods: To address the limitations of existing approaches, we propose MediHerb, a multi-modal enhanced framework for disease inference via herbal knowledge. MediHerb unifies five complementary modalities: molecular sequences, fingerprints, physicochemical properties, graphical prescription representations, and the description of TCM prescriptions into a shared latent space. An attention-based fusion mechanism aligns the semantics across molecular, herbal, and diagnostic levels, enabling multi-granularity reasoning. To further promote accessibility, a lightweight graphical interface has been developed to support interaction with both the model and open datasets.
Results: Experimental results on benchmark datasets demonstrate that MediHerb substantially outperforms existing baselines in herb–disease inference. Beyond predictive accuracy, the learned embeddings and model attention patterns reveal meaningful biological and pharmacological insights, confirming that MediHerb captures the mechanistic underpinnings of herb–disease associations.
Conclusion: MediHerb highlights the potential of knowledge-enhanced multi-modal fusion to bridge molecular, herbal, and clinical semantics, offering a more interpretable and holistic approach to understanding TCM prescriptions.
Less -
Xiaoyi Liu, ... Jijun Tang
-
DOI: https://doi.org/10.70401/cbm.2025.0003 - November 25, 2025
Drug-target affinity prediction based on multi-source information and graph convolutional network
-
Aims: Drug-target affinity (DTA) prediction is crucial for drug discovery and repositioning. However, existing deep learning-based methods often overlook the synergy between the topological structure of DTA networks and the multimodal features ...
MoreAims: Drug-target affinity (DTA) prediction is crucial for drug discovery and repositioning. However, existing deep learning-based methods often overlook the synergy between the topological structure of DTA networks and the multimodal features of drugs and targets themselves.
Methods: This study proposes a new method, MIGDTA, a DTA prediction method based on multi-source information and graph convolutional network (GCN), which enhances prediction accuracy by integrating local features with global interaction information. MIGDTA first constructs a drug molecular graph, a target protein graph, and a DTA network, while computing molecular fingerprints and protein descriptors. Subsequently, it employs a graph isomorphism network to learn graph features, a GCN to capture network features, and a multilayer prceptron to encode biological features. Then, it refines heterogeneous network and graph features iteratively through the GCN, and finally concatenates the fused features with biological features for affinity prediction.
Results: Comparative experiments on benchmark datasets demonstrate that MIGDTA significantly outperforms existing methods. On the Davis dataset, compared to the best baseline method, MIGDTA reduces mean squared error (MSE) to 0.185, increases CI by 0.006, and improves
by 5%. Similar enhancements were observed on the KIBA dataset, where MIGDTA achieves an MSE of 0.130, along with 0.002 and 1% gains in CI and , respectively. Conclusion: Feature ablation studies verify the core role of graph features in modeling local structures and network features in capturing global topology, along with the supplementary importance of biological features. Comparative analyses of feature integration approaches confirm the effectiveness of the feature refinement module in fusing multimodal features and enhancing model discriminability.
Less -
Xiujuan Lei, ... Yuchen Zhang
-
DOI: https://doi.org/10.70401/cbm.2026.0007 - January 19, 2026
A bi-directional LSTM architecture enhanced with channel attention for seizure prediction
-
Aims: Neural networks capable of capturing temporal dependencies in electroencephalogram (EEG) signals hold considerable potential for seizure prediction by modeling the progressive evolution of preictal EEG changes. However, redundant or less ...
MoreAims: Neural networks capable of capturing temporal dependencies in electroencephalogram (EEG) signals hold considerable potential for seizure prediction by modeling the progressive evolution of preictal EEG changes. However, redundant or less informative temporal features may obscure critical preictal patterns, limiting seizure prediction performance. To address this, we developed a neural architecture that effectively leverages informative temporal features to enhance seizure prediction capability.
Methods: We designed a bidirectional long short-term memory (BiLSTM) network enhanced with a channel attention mechanism, termed Attention-BiLSTM, which adaptively emphasizes informative temporal features while reducing information redundancy. We further analyze the model’s attention weights and feature distributions to provide interpretable insights into its decision-making process.
Results: Evaluation on the CHB-MIT scalp EEG dataset demonstrates that Attention-BiLSTM achieves significant performance improvements over the baseline BiLSTM, with an average accuracy of 94.77%, sensitivity of 94.58%, specificity of 94.97%, and an area under the curve of 98.38%. Furthermore, visualization results indicate that the proposed model progressively enhances feature discriminability and directs attention to the most relevant temporal features for seizure prediction.
Conclusion: The proposed Attention-BiLSTM achieves improved performance and interpretability, offering valuable insights to support future development of scalable and generalizable seizure prediction systems.
Less -
Haiqing Yu, ... Dong Ming
-
DOI: https://doi.org/10.70401/cbm.2026.0010 - February 05, 2026
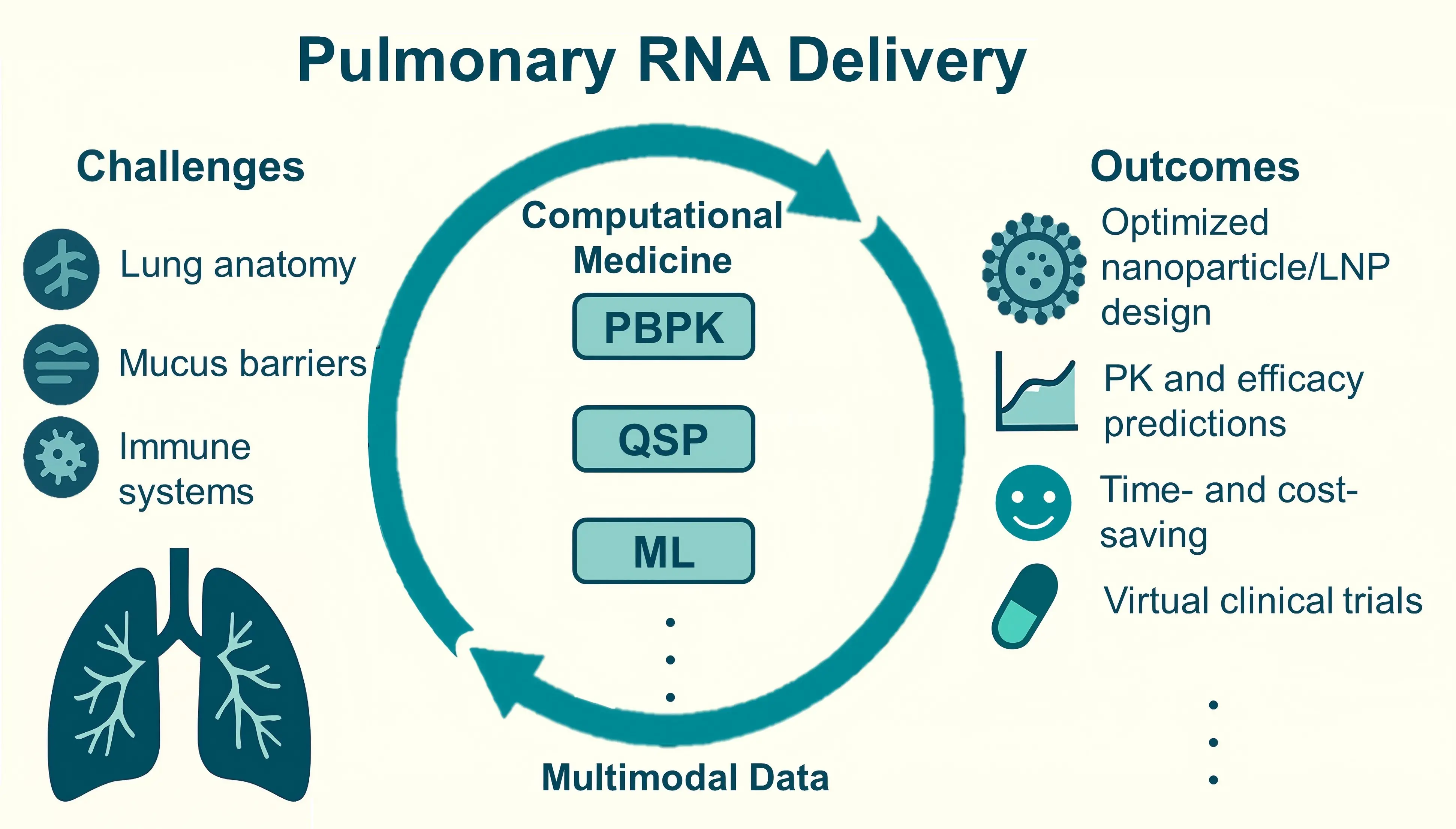
Computational approach to pulmonary delivery of therapeutical RNAs
-
Targeted delivery of RNA-based therapeutics to the lungs remains a substantial challenge due to the unique anatomy of lung tissue and its complex immune barriers. In recent years, the convergence of physiologically based pharmacokinetic (PBPK) models, quantitative ...
MoreTargeted delivery of RNA-based therapeutics to the lungs remains a substantial challenge due to the unique anatomy of lung tissue and its complex immune barriers. In recent years, the convergence of physiologically based pharmacokinetic (PBPK) models, quantitative systems pharmacology approaches, and machine learning algorithms has led to the development of computational medicine frameworks, providing intelligent tools for addressing the aforementioned challenges in efficient pulmonary delivery. By integrating experimental data with predictive computational models, these approaches have advanced the development of RNA therapies for pulmonary diseases. The deep integration of multimodal data is expected to further accelerate drug discovery and clinical translation. In this review, we systematically summarize these computational approaches in designing and optimizing pulmonary RNA delivery systems. We particularly highlight the mechanism-based rational design of RNA therapies through simulations and predictions of biodistribution, cellular targeting, and intracellular transport processes.
Less -
Xianan Li, ... Pu Chen
-
DOI: https://doi.org/10.70401/cbm.2025.0004 - November 28, 2025
A comprehensive review on neuropeptides: databases and computational tools
-
Neuropeptides are crucial signaling molecules that regulate diverse physiological processes spanning growth, social behavior, learning, memory, metabolism, homeostasis, reproduction, and neural differentiation across both nervous and peripheral ...
MoreNeuropeptides are crucial signaling molecules that regulate diverse physiological processes spanning growth, social behavior, learning, memory, metabolism, homeostasis, reproduction, and neural differentiation across both nervous and peripheral systems. Dysregulation of neuropeptides signaling is closely linked to various pathological conditions, such as neurological disorders, metabolic diseases, cardiovascular conditions, and even cancer, positioning them as potential therapeutic agents or targets for intervention. In recent years, research into neuropeptides has accelerated, with vast amounts of data continuously accumulating in multiple databases. However, the study of neuropeptides is often impeded by the need for extensive and time-consuming experimental investigations. As a result, computational tools have become essential for the rapid, large-scale identification of neuropeptides. This review systematically discusses neuropeptide-related databases and computational tools. These databases organize extensive data on neuropeptide sequences, structures, and functions. Among these, NeuroPep2.0, with 11,417 neuropeptide entries, is currently the most widely used dataset for neuropeptide prediction. Additionally, this review explores the application of computational approaches in neuropeptide prediction. While early methods predominantly relied on homologous sequence alignment and biochemical feature statistics, recent advances in machine learning have significantly enhanced prediction accuracy and efficiency. Tools such as NeuroPred-PLM and DeepNeuropePred, developed by our research group using protein language models, have substantially improved prediction performance. In conclusion, this review provides a comprehensive overview of current neuropeptide databases and computational tools, offering researchers a thorough survey of available resources and analytical methods, and emphasizing the necessity of continuous optimization to advance neuropeptide research and its therapeutic applications.
Less -
Wei Xu, ... Yan Wang
-
DOI: https://doi.org/10.70401/cbm.2025.0001 - April 10, 2025
MediHerb: A multi-modal enhanced framework for disease inference via herbal knowledge
-
Aims: Development of robust and effective methods for uncovering herb interactions and constructing herb–disease associations requires the integration of diverse biological and medical information. A key challenge in Traditional Chinese Medicine ...
MoreAims: Development of robust and effective methods for uncovering herb interactions and constructing herb–disease associations requires the integration of diverse biological and medical information. A key challenge in Traditional Chinese Medicine (TCM) research is to robustly uncover herb interactions and construct reliable herb–disease associations. This task requires handling the inherently high-dimensional, multi-label, and cross-domain nature of prescription data. Existing approaches provide limited representation capacity and insufficient integration of biomedical knowledge, restricting their ability to capture the complex semantics underlying the relationships between herbs and diseases.
Methods: To address the limitations of existing approaches, we propose MediHerb, a multi-modal enhanced framework for disease inference via herbal knowledge. MediHerb unifies five complementary modalities: molecular sequences, fingerprints, physicochemical properties, graphical prescription representations, and the description of TCM prescriptions into a shared latent space. An attention-based fusion mechanism aligns the semantics across molecular, herbal, and diagnostic levels, enabling multi-granularity reasoning. To further promote accessibility, a lightweight graphical interface has been developed to support interaction with both the model and open datasets.
Results: Experimental results on benchmark datasets demonstrate that MediHerb substantially outperforms existing baselines in herb–disease inference. Beyond predictive accuracy, the learned embeddings and model attention patterns reveal meaningful biological and pharmacological insights, confirming that MediHerb captures the mechanistic underpinnings of herb–disease associations.
Conclusion: MediHerb highlights the potential of knowledge-enhanced multi-modal fusion to bridge molecular, herbal, and clinical semantics, offering a more interpretable and holistic approach to understanding TCM prescriptions.
Less -
Xiaoyi Liu, ... Jijun Tang
-
DOI: https://doi.org/10.70401/cbm.2025.0003 - November 25, 2025
Drug-target affinity prediction based on multi-source information and graph convolutional network
-
Aims: Drug-target affinity (DTA) prediction is crucial for drug discovery and repositioning. However, existing deep learning-based methods often overlook the synergy between the topological structure of DTA networks and the multimodal features ...
MoreAims: Drug-target affinity (DTA) prediction is crucial for drug discovery and repositioning. However, existing deep learning-based methods often overlook the synergy between the topological structure of DTA networks and the multimodal features of drugs and targets themselves.
Methods: This study proposes a new method, MIGDTA, a DTA prediction method based on multi-source information and graph convolutional network (GCN), which enhances prediction accuracy by integrating local features with global interaction information. MIGDTA first constructs a drug molecular graph, a target protein graph, and a DTA network, while computing molecular fingerprints and protein descriptors. Subsequently, it employs a graph isomorphism network to learn graph features, a GCN to capture network features, and a multilayer prceptron to encode biological features. Then, it refines heterogeneous network and graph features iteratively through the GCN, and finally concatenates the fused features with biological features for affinity prediction.
Results: Comparative experiments on benchmark datasets demonstrate that MIGDTA significantly outperforms existing methods. On the Davis dataset, compared to the best baseline method, MIGDTA reduces mean squared error (MSE) to 0.185, increases CI by 0.006, and improves
by 5%. Similar enhancements were observed on the KIBA dataset, where MIGDTA achieves an MSE of 0.130, along with 0.002 and 1% gains in CI and , respectively. Conclusion: Feature ablation studies verify the core role of graph features in modeling local structures and network features in capturing global topology, along with the supplementary importance of biological features. Comparative analyses of feature integration approaches confirm the effectiveness of the feature refinement module in fusing multimodal features and enhancing model discriminability.
Less -
Xiujuan Lei, ... Yuchen Zhang
-
DOI: https://doi.org/10.70401/cbm.2026.0007 - January 19, 2026
iCDG-MOHGAT: Identification of cancer driver gene using multi-omics data and heterogeneous graph attention network
-
Aims: Driver mutations are crucial factors in the occurrence and development of cancer. Identifying cancer-related driver genes is of great significance for understanding the mechanisms of cancer initiation, prevention, and treatment. With the ...
MoreAims: Driver mutations are crucial factors in the occurrence and development of cancer. Identifying cancer-related driver genes is of great significance for understanding the mechanisms of cancer initiation, prevention, and treatment. With the continuous accumulation of cancer data, how to effectively utilize these data for the identification of cancer driver genes has become a major challenge in the field of cancer biology.
Methods: We propose a novel computational model called iCDG-MOHGAT. This model integrates multi-omics pan-cancer data (such as mutations, DNA methylation, etc.), multi-dimensional gene networks, and disease semantic similarity networks to identify cancer driver genes. We first construct multi-dimensional gene networks using various types of gene correlation information (protein-protein interaction, gene sequence similarity, etc.) and establish disease semantic similarity networks for relevant cancers. Due to the complexity of node and edge types, we utilize a heterogeneous graph attention network to learn and extract features from the multi-dimensional gene networks and disease semantic similarity networks. We also incorporate a fusion learning module to effectively integrate features from different dimensions. Finally, we optimize the random forest classifier using the sparrow algorithm for the task of predicting cancer driver genes.
Results: Experimental results demonstrate that iCDG-MOHGAT outperforms many state-of-the-art models in terms of AUPR and AUROC. In the final prediction results, 91% of the predicted new driver genes have at least one supporting evidence of being cancer genes. In the laboratory, this model can serve as an effective tool for identifying cancer driver genes.
Conclusion: We have introduced a novel computational model named iCDG-MOHGAT, which precisely identifies cancer driver genes by integrating multi-omics pan-cancer data and intricate multidimensional gene networks, coupled with disease semantic similarity networks. Experimental results demonstrate that iCDG-MOHGAT outperforms many state-of-the-art models in terms of AUPR and AUROC. In the final prediction results, 91% of the predicted genes have supporting evidence. In the laboratory, this model can serve as an effective tool for identifying cancer driver genes.
Less -
Lin Yuan, Jiawang Zhao
-
DOI: https://doi.org/10.70401/cbm.2026.0008 - February 03, 2026
A bi-directional LSTM architecture enhanced with channel attention for seizure prediction
-
Aims: Neural networks capable of capturing temporal dependencies in electroencephalogram (EEG) signals hold considerable potential for seizure prediction by modeling the progressive evolution of preictal EEG changes. However, redundant or less ...
MoreAims: Neural networks capable of capturing temporal dependencies in electroencephalogram (EEG) signals hold considerable potential for seizure prediction by modeling the progressive evolution of preictal EEG changes. However, redundant or less informative temporal features may obscure critical preictal patterns, limiting seizure prediction performance. To address this, we developed a neural architecture that effectively leverages informative temporal features to enhance seizure prediction capability.
Methods: We designed a bidirectional long short-term memory (BiLSTM) network enhanced with a channel attention mechanism, termed Attention-BiLSTM, which adaptively emphasizes informative temporal features while reducing information redundancy. We further analyze the model’s attention weights and feature distributions to provide interpretable insights into its decision-making process.
Results: Evaluation on the CHB-MIT scalp EEG dataset demonstrates that Attention-BiLSTM achieves significant performance improvements over the baseline BiLSTM, with an average accuracy of 94.77%, sensitivity of 94.58%, specificity of 94.97%, and an area under the curve of 98.38%. Furthermore, visualization results indicate that the proposed model progressively enhances feature discriminability and directs attention to the most relevant temporal features for seizure prediction.
Conclusion: The proposed Attention-BiLSTM achieves improved performance and interpretability, offering valuable insights to support future development of scalable and generalizable seizure prediction systems.
Less -
Haiqing Yu, ... Dong Ming
-
DOI: https://doi.org/10.70401/cbm.2026.0010 - February 05, 2026
Frontier Forums
Special Issues
AI for Biomedicine: Models, Applications, and Challenges
-
Submission Deadline: 31 Aug 2026
-
Published articles: 1
-
Submission Deadline: 31 Aug 2026
-
Published articles: 0