Jessica K. Tyler, Weill Cornell Medicine, Department of Pathology and Laboratory Medicine, New York, NY 10065, USA. E-mail: jet2021@med.cornell.edu
Abstract
The nucleolus, the largest membraneless organelle in the cell, is a biomolecular condensate that houses ribosomal DNA (rDNA), facilitates ribosomal subunit assembly, and serves as a dynamic reservoir for numerous unrelated proteins. Aging across eukaryotic species is accompanied by nucleolar expansion, raising the question of whether it is a correlate of aging or a driver of cellular aging. Recent studies suggest that nucleolar expansion may drive aging and this may result from age-associated changes in the biophysical properties of the nucleolus. Emerging evidence points to age-driven biophysical changes in the nucleolar condensate, including shifts in size, dynamics, and viscoelasticity, which may occur gradually or through transitions from a liquid-like state to denser gel-like, and in some contexts amyloid-like, assemblies. These transitions remodel two core condensate properties: compartmentalization and partitioning, with consequences for ribosome biogenesis and rDNA stability. Here, we review recent literature on age-driven changes in nucleolar condensation and discuss how these changes may influence nucleolar function and longevity.
Keywords
1. Introduction
Most interventions known to enhance longevity in mammals are closely linked to cellular growth and metabolism (extensively reviewed in previous studies[1,2]). For instance, dietary restriction and reduced insulin/IGF-1 signaling extend lifespan by suppressing growth-promoting pathways and enhancing stress resistance[1,3]. Similarly, inhibition of mTOR/Tor1, either genetically or pharmacologically, extends lifespan from mammals to yeast by activating autophagy, and concurrently reduces protein synthesis[4]. Activation of AMPK, a central energy sensor that promotes catabolic over anabolic processes, also enhances longevity by improving metabolic efficiency and stress tolerance[5]. Together, these findings suggest that lifespan extension can be achieved by reprogramming cellular priorities from biosynthesis and proliferation toward stress resilience and reduced anabolic demand. Ribosome biogenesis is a major consumer of biosynthetic resources, and ribosome abundance itself determines translational capacity and consequently, metabolic flux.
The nucleolus, where ribosomal DNA (rDNA) is transcribed into ribosomal RNA (rRNA) and further processed into ribosomal subunits, sits at the heart of this growth–metabolism axis[6]. Historically, nucleolar expansion was among the first recognized cellular phenotypes of cellular aging[7], although the underlying reason was unclear. Coherently, nucleolar size decreases in response to several lifespan-extending regimens, such as reduced insulin/IGF signaling, mTOR inhibition and dietary restriction, providing a common morphological signature of pro-longevity physiology[8]. Yet it remained unresolved whether the small nucleolar size observed under pro-longevity regimens merely mirrors metabolic reprogramming or can actively drive anti-aging. Our recent work supports the latter view[9], where the increase in nucleolus size and accompanied rDNA instability during aging act as a countdown to cell death, i.e., as a mortality timer. Nucleolar size has long served as a proxy for ribosome biogenesis output, where larger nucleoli correlate with greater ribosome biogenesis[8]. As phase-separated condensates, nucleoli exhibit changes in size or number that often reflect underlying shifts in biophysical properties, such as liquidity, viscosity, or fusion behavior[10,11], which can influence nucleolar function during aging[8,9]. In this review, we discuss how these nucleolar variables change with age and whether they, in turn, influence longevity.
Historically, the nucleolus has been recognized since the late 18th century: Felice Fontana sketched it in 1781, well before ‘organelle’ entered the cell-biology lexicon. Only recently, however, have its multidimensional functions begun to be elucidated. Unlike membrane-bound organelles, the nucleolus lacks a delimiting membrane, forms through condensation of nucleic acids and proteins, and scaffolds into coexisting subcompartments, providing compartmentalization of nucleolar processes. Selective partitioning of proteins and RNAs across these subcompartments creates a specialized chemical and physical environment optimized not only for ribosomal subunit assembly but also for housing and maintenance of the fragile, repetitive rDNA, which encodes the RNA components of ribosomes[12]. Notably, the nucleolus also selectively recruits proteins not dedicated to ribosome biogenesis or rDNA related
2. The Nucleolus is A Biomolecular Condensate
From the earliest light micrographs, reviewed by Thomas H. Montgomery in 1898, the nucleolus was described as a darker, distinct body within the nucleus, providing an early indication of nuclear compartmentalization[14]. Classic cytochemistry and extraction experiments then suggested a matrix- or gel-like structure, as removal of RNA by ribonuclease treatment and partial pepsin digestion left a stable fibrillar meshwork of proteins and DNA[15]. However, a landmark study in Xenopus laevis eggs from Tony Hyman’s lab provided the first direct biophysical evidence that nucleoli behave as phase-separated liquid droplets that fuse and flow with viscosity and surface tension[10]. Importantly, the nucleolus is not a homogeneous liquid structure but contains coexisting
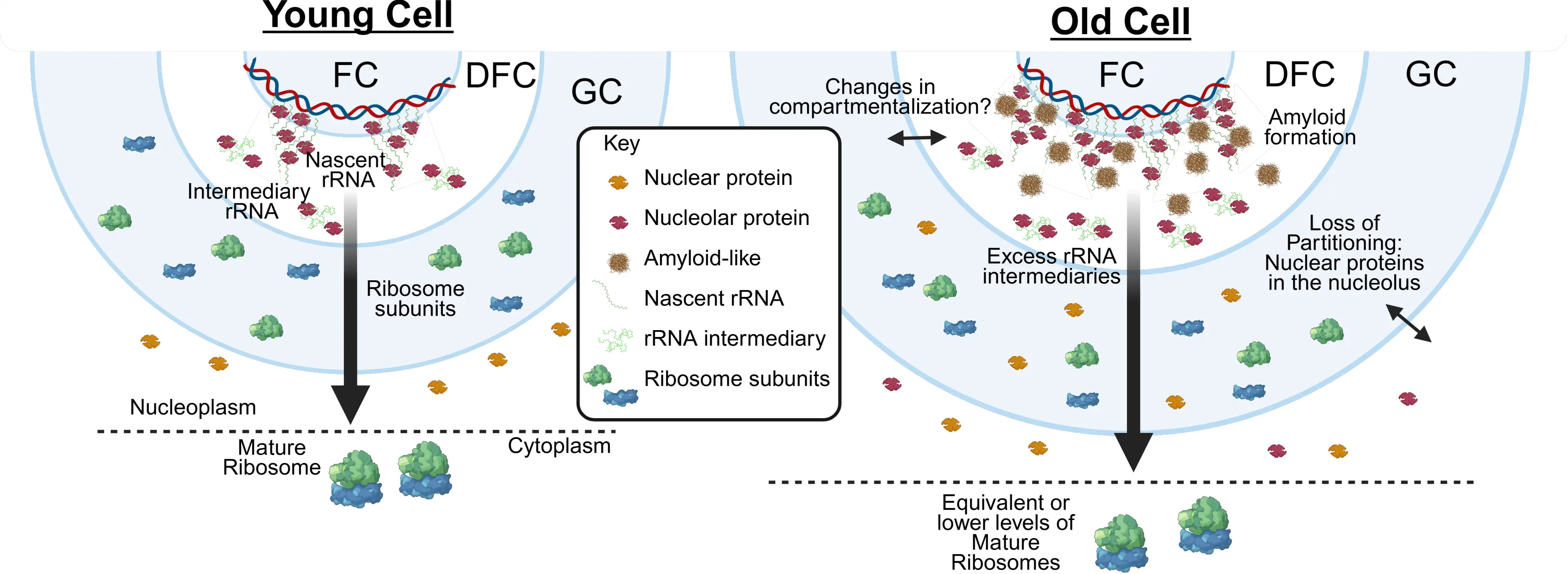
Figure 1. Model for age-associated changes in nucleolar organization. In old cells, rDNA transcription increases while ribosome biogenesis remains unchanged, leading to an accumulation of rRNA intermediates. This imbalance may alter the biophysical properties of the nucleolus, including the appearance of amyloid-like structures in the FC/DFC and reduced partitioning fidelity, allowing nuclear proteins to aberrantly accumulate within the nucleolus. These changes may also affect nucleolar compartmentalization during aging, although this possibility remains to be demonstrated. rDNA: ribosomal DNA; rRNA: ribosomal RNA; FC: fibrillar center; DFC: dense fibrillar component; GC: granular component.
The nucleolar subcompartments form through self-assembly into emulsion-type subphases, driven by the biochemical properties of their constituent proteins. In vitro reconstitution experiments reveal that recombinant nucleolar scaffold proteins such as nucleophosmin (NPM1, found in the GC) and fibrillarin (FIB1, found in the DFC), can independently undergo phase separation, recreating distinct nucleolar subcompartments when mixed in vitro[16-18]. Interestingly, siRNA knock-down of NPM1[19], FIB1[20] and other nucleolar proteins alters nucleolar morphology and affects ribosome biogenesis, but does not prevent nucleolar formation, indicating that such formation relies on redundant or overlapping scaffolding mechanisms. Both NPM1 and FIB1 contain
3. Nucleolar Expansion During Aging
In the late 1970s, Bemiller & Lee observed that nucleolar size increases dramatically in old senescent cultured human fibroblasts, with multiple smaller nucleoli often coalescing into a single, enlarged structure, a phenomenon subsequently confirmed in other aged organisms, including yeast and worms[7,25,26] (Figure 1). In yeast, deletion of SGS1, the homolog of the human WRN gene whose mutation causes Werner’s syndrome, a premature aging disorder, results in abrupt nucleolar expansion and fragmentation during aging, accompanied by a reduction in replicative lifespan (i.e., the number of cell divisions in a cell’s lifespan)[27]. Consistently, the nucleoli from fibroblasts isolated from humans of different ages show increased size with age, while fibroblasts from young patients with Hutchinson Gilmore Progeria Syndrome, another premature aging disorder, display enlarged nucleoli[26]. Supporting the idea that nucleolar enlargement can reflect a pathogenic nucleolar stress state, systemic induction of nucleolar stress in mice through expression of arginine-rich poly(PR) peptides increases fibrillarin-positive nucleolar area across tissues and accelerates organismal aging, leading to premature death[28]. Conversely, small nucleolar size is induced by numerous pro-longevity regimens[29]. Reduction of nucleolar size is also observed during gametogenesis-mediated cellular rejuvenation in yeast[30], and during C. elegans lifespan extension caused by RNAi depletion of the nucleolar protein fibrillarin[8]. Strikingly, in C. elegans, nucleolar size in hypodermal cells measured on the first day of adulthood inversely correlates with remaining lifespan, further strengthening the negative correlation between nucleolar size and longevity across taxa[8].
In young cells, nucleolar size is closely linked to rDNA transcription[31,32]. Elevated rDNA transcription drives increased ribosomal subunit biogenesis, accompanied by nucleolar enlargement. Size is a highly dynamic feature of biomolecular condensates[18,24], and modulation of the nucleolar size in response to rRNA synthesis could help maintain constant molecular crowding and biophysical properties to allow efficient ribosome assembly. Indeed, in aged human fibroblasts and fibroblasts from young progeria patients, nucleolar expansion is associated with increased ribosome biogenesis and an overall increase in protein synthesis[26]. However, this is not the case in other cell types and organisms because in yeast, age-driven nucleolar expansion occurs without increased ribosome biogenesis[8,9,33]. In multiple human cell types, including neurons, hepatocytes, and muscle cells, aging is accompanied by a modest decline in ribosome abundance and translational efficiency[34-36]. This raises the possibility that age-associated dissociation between nucleolar size and ribosome biogenesis may contribute to the detrimental impact of nucleolar expansion on longevity (Figure 1). A non-aging pathological example of such imbalance between nucleolar size and ribosome biogenesis is observed in spinal muscular atrophy (SMA) motor neurons, where nucleoli become markedly enlarged and the FC subcompartment expands while decreasing in number. Notably, SMA neurons also show altered rRNA homeostasis, including accumulation of rRNA species, supporting the idea that nucleolar hypertrophy can reflect stalled or dysregulated ribosome production rather than elevated biosynthetic
During aging, it is not yet clear whether the size of all nucleolar subcompartments is enlarged or whether only specific subcompartments are affected. Clarifying this distinction could provide important insight into the molecular reason for their enlargement. For example, preferential enlargement of the GC subcompartment may indicate impaired nucleolar export of
To distinguish whether nuclear expansion is a cause or a consequence of aging, we engineered yeast to express dCas9 fused to the inner nuclear membrane protein Heh1, guided by RNAs targeting the rDNA[9]. This rDNA-tethering system delayed expansion of the yeast nucleolus during aging, resulting in lifespan extension[9]. Even in wild-type cells, the yeast that expanded their nucleolus later during aging, lived longer[9]. Thus, while nucleolar expansion occurs as a consequence of aging, it clearly limits cellular longevity. Furthermore, live cell imaging of the GFP-tagged yeast nucleolus throughout the lifespan of yeast trapped in microfluidic devices revealed an unexpected finding. Although nucleolar expansion progresses gradually during aging, once a critical size termed the Nucleolar Size Threshold is reached, subsequent expansion accelerates markedly[9]. Strikingly, after the nucleolar size threshold was surpassed, the cells divided on average only five more times before death. Together, these findings suggest that nucleolar expansion, upon reaching a certain threshold, marks a transition to a new biophysical state that functions as a mortality timer during aging
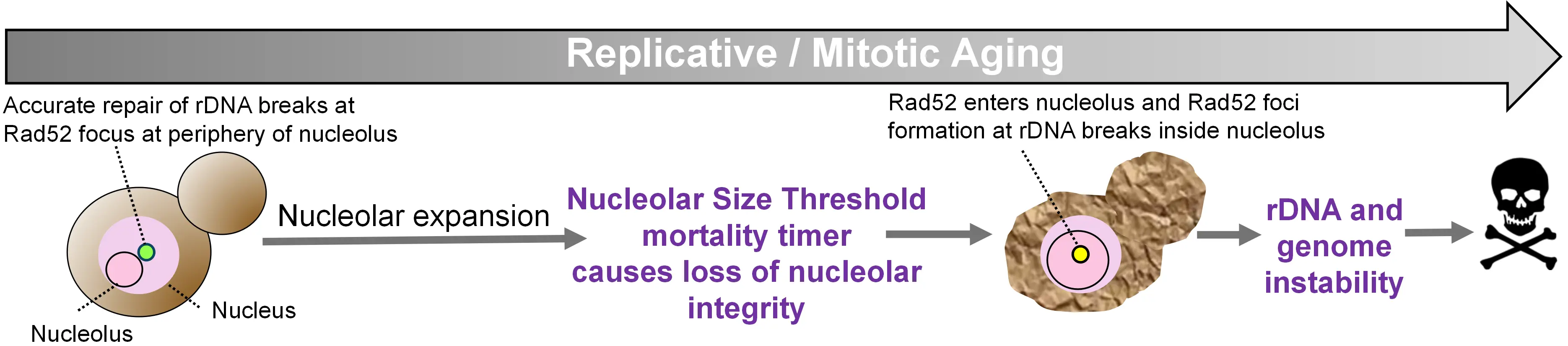

Figure 2. Nucleolar size threshold as an irreversible transition during yeast replicative aging that causes rDNA instability and ends the lifespan. In young cells, and for most of the replicative lifespan, the nucleolus remains small and repair of rDNA DSBs is restricted to the nucleolar periphery. As cells age, nucleolar expansion accelerates abruptly upon approaching the NST. Once the NST is exceeded, nucleolar integrity is lost and Rad52 enters the nucleolus, forming repair foci at rDNA breaks within the nucleolar compartment, triggering inaccurate rDNA recombination, associated global genome instability and imminent cell death, consistent with the NST acting as a mortality timer. rDNA: ribosomal DNA; DSBs: DNA double-strand breaks; NST: nucleolar size threshold.
4. rDNA Instability and Age-Driven Nucleolar Expansion
DNA double-strand breaks (DSBs) within the rDNA occur naturally across organisms from yeast to humans, arising from a roadblock mechanism that prevents collisions between the DNA replication and transcription machinery[42]. However, rDNA instability greatly increases with age, as observed in yeast[43,44], Drosophila[45,46], mice[47] and human aging[48], as well as in age-related disease[49] and progeroid contexts[50]. rDNA instability can be easily detected during yeast aging[44], due to the presence of a single rDNA locus that can be excised from the genome and resolved by pulsed-field electrophoresis to detect gains and losses of rDNA repeats, and by the accumulation of extrachromosomal rDNA circles (ERC) evicted from the genome as a consequence of intrachromosomal rDNA recombination[43]. In addition, the rDNA locus in aged yeast was identified as a hotspot of gammaH2A (the yeast equivalent of the gammaH2AX marker of DNA DSBs) accumulation in ChIP-seq analyses, a frequent translocation partner in age-induced chromosomal translocations, and a site of recurrent amplification of the chromosomal region distal to the rDNA locus[51]. By contrast, detection of rDNA instability during aging in mammals remains technically challenging because the rDNA repeats are distributed across multiple acrocentric chromosomes, are highly homogeneous, and are poorly resolved by standard genomic and cytological
We recently uncovered that age-driven nucleolar expansion in yeast triggers rDNA instability due to mislocalization of the DNA repair machinery (Figure 2). DNA DSBs within the rDNA are normally repaired by homologous recombination at the nucleolar periphery to favor use of the corresponding undamaged rDNA repeat from the sister chromatid as the homology template. This deters aberrant recombination using one of the ~150 other rDNA repeats as the homology template, which may occur if homologous recombination took place within the nucleolus in yeast[56]. Homologous recombination of rDNA breaks is limited to the nucleolar periphery as the DNA DSB relocate to this region[56]. While in mammalian cells, rDNA DSBs trigger the formation of a nucleolar cap at the periphery, a specialized repair compartment comprising rDNA with FC/DFC material[57]. In yeast, homologous recombinational repair of rDNA DSBs is restricted to the nucleolar periphery because Rad52, the yeast functional equivalent of mammalian Brca2, is actively expelled from the nucleolus, allowing repair to proceed only after the rDNA DSBs exit the nucleolus[56]. Unexpectedly, we observed that in old yeast with nucleoli exceeding the nucleolar size threshold, Rad52-GFP foci appeared within the nucleolus rather than at the periphery, triggering rDNA hyperrecombination and imminent death[9] (Figure 2).
In old yeast lacking Rad52 foci, free Rad52 and another normally nucleolar-excluded protein were observed to enter the expanded nucleoli[9]. Furthermore, a large nucleolus is sufficient for Rad52 entry, given that expansion of nucleoli in young yeast, induced by treatment with the Sirtuin inhibitor resveratrol, similarly resulted in Rad52-GFP foci within the nucleoli[9]. Notably, an engineered system that delays yeast nucleolar expansion during aging resulted in reduced rDNA instability, as detected by ERC formation and lifespan extension[9], highlighting the relationship between rDNA instability and nucleolar size. Interestingly, recent work in
Importantly, age-induced rDNA instability also promotes global genome instability. This was evident in the reduced chromosomal fragmentation of yeast deleted for FOB1, the roadblock protein that induces the rDNA breaks[44]. Furthermore, engineering the yeast nucleolus to remain smaller during aging both reduced and delayed the appearance of Rad52-GFP foci, reflecting sites of homologous recombinational repair[9]. Exactly why rDNA instability during aging induces genome-wide instability is not clear, but it may reflect competition for limited pools of DNA repair factors[44]. Regardless, these findings underscore that nucleolar size has implications beyond rDNA instability alone, presumably contributing to cellular death. What is driving the changes in the nucleolar condensate that correlate with, and potentially cause, nucleolar expansion, leading to age-induced rDNA instability and end of lifespan?
5. Changes in the Condensation Properties of the Nucleolus
Biomolecular condensates span a continuum of material states; their components can be diluted in solution, or phase-separate into liquid, gel-like, or solid amyloid-like structures. Small environmental or chemical changes, such as variations in pH, ions, RNA/protein stoichiometry and post-translational modifications, can shift them along this spectrum in a highly dynamic
6. Age-Driven Changes in Nucleolar Condensation
The idea that aberrant phase transitions of condensates could be a driver of cellular aging has been hypothesized[65], but direct experimental evidence for this mechanism has recently begun to emerge. Across species, nucleolar aging has been associated with densification, i.e., transitioning from a liquid-like state to a gel and in some cases, an amyloid-like state. In intestinal epithelial cells of C. elegans, FIB1-GFP (fibrillarin, localized at the rDNA transcription site DFC) shows slower FRAP recovery in old adults than in larvae, indicating reduced molecular exchange within the rRNA-processing layer and a shift toward a more gel-like nucleolar state[16]. Notably, in vitro reconstitution of fibrillarin-rich droplets reveals time-dependent “maturation”, slower fusion and rounding and diminished internal rearrangements, coherent with the in vivo slower FRAP recovery[16]. Yeast replicative aging experiments, which model the functional decline accompanying successive cell divisions, show that expanded nucleolus beyond a critical size threshold becomes resistant to 1,6 hexanediol dissolution[9]. Importantly, resistance to 1,6 hexanediol (as well as reduced FRAP recovery) indicates decreased condensate dynamics, but does not constitute definitive evidence for cross-β amyloid fibril formation[66].
In human cell systems, cellular senescence provides an experimentally tractable model for cellular aging, i.e., a stable cell growth arrest after a finite number of divisions[68], triggered by telomere attrition and/or DNA-damage. Like yeast replicative aging, it culminates in loss of proliferative capacity and features nucleolar expansion[7,25,26,68]. The accumulation of senescent cells is a hallmark of aging[69]. Several cell types undergo senescence induced by a group of agents that reduce genome integrity by causing DNA lesions/replication stress, and by agents that disrupt gene expression programs (rDNA/Pol I, Pol II, epigenetic modifiers); additional drug classes (CDK4/6 inhibitors, proteostasis stressors) also drive senescence via non-genotoxic routes[70]. Importantly, senescence induced by Pol I inhibition[71], CDK4 inhibition[72], oxidative stress[73] and DNA damage[74], although not identical to replicative senescence[70], conserves the expanded nucleolus phenotype[73,74]. Under multiple in vitro stress-induced senescence models, the density of the DFC subcompartment of the nucleolus increases, as measured by FIB1-GFP FRAP[73], similarly to that observed in C. elegans[16]. Moreover, super resolution imaging shows the appearance of fibrils stained by Congo red strongly suggesting the appearance of amyloid-like features in the nucleolus[73]. Similarly, another study using label-free vibrational spectroscopy in binucleation-driven senescence identified β-sheet–like, aggregation-prone signatures within nucleoli, providing molecular evidence for nucleolar solidification[74].
Changes in the phase behavior of proteins with low-complexity domains have been extensively studied in the context of neurodegeneration and age-associated neurological disorders, particularly with respect to the formation of amyloid-like aggregates (reviewed in the work[75]). A recent study reported that amyloid-positive nucleolar “cavities” are prevalent in postmortem human neurons from aged brains, including neurologically healthy individuals and patients with neurodegenerative diseases. These cavities are intranucleolar inclusions that exclude canonical nucleolar markers such as NPM1, FIB1 and PolI subunit RPA194, but stain positively with Congo red, consistent with amyloid-like material[76]. While these structures were common in aged neurons across conditions, neurodegenerative disease was associated with the selective sequestration of specific aggregation-prone proteins within the nucleolus, including phosphorylated tau and TDP-43. Notably, rDNA instability has previously been linked to neurodegenerative diseases[49], raising the possibility that age-associated changes in nucleolar condensation may contribute to genomic instability at rDNA loci, although a causal relationship remains to be established. Collectively, these studies suggest that across taxa
What drives the denser nucleolar phenotype during aging remains unclear; however, one possible explanation could be a reported increase in PolI-mediated rRNA production[26,33] that is not accompanied by increases in ribosome biogenesis[9,33], creating “traffic” in the unidirectional highway of ribosome biogenesis within the nucleolus. Indeed, perturbations that stall early pre-rRNA processing slow FRAP recoveries, indicating increased density/crowding at the inner core of the nucleolus[22], whereas inhibition of PolI leads to faster FRAP recoveries[77]. Collectively, these findings suggest a conserved biophysical alteration in aging cells where stress-induced senescence and dysregulated rRNA flux coincide with the transformation of the nucleolus into a dense, aggregation-prone compartment, potentially disrupting the spatial organization and partitioning of nucleolar proteins (Figure 1). Whether this phenotype is fully recapitulated during replication-induced senescence remains to be determined.
7. Nucleolar Partitioning and Compartmentalization
Condensation (phase separation) propensity and partitioning are not identical, although they often correlate because both depend on the same multivalent interactions[60,78]. Partitioning refers to the thermodynamic preference to reside in the nucleolar network (partition coefficient: nucleolus/nucleoplasm); phase separation describes the emergent formation of a dense biomolecular phase coexisting with a surrounding dilute phase[11,60]. Thus, a mutation can in principle lower phase separation while maintaining or even raising partitioning (e.g., by strengthening specific heterotypic binding), or reduce partitioning without abolishing phase separation (e.g., if stoichiometry becomes limiting)[17,79]. As mentioned above, nucleolar condensation relies on multiple overlapping scaffolding mechanisms. Mutations targeting the acidic A-tracts and basic B-tracts of NPM1, charged sequence motifs within its intrinsically disordered region that mediate multivalent interactions with rRNA and protein partners[17], provide a concrete example, at least for NPM1, for assessing the relationship between condensation and partitioning of the nucleolus. In vitro/in vivo comparisons show clear concordance for this set of NPM1 mutants. NMP1 mutA3 lowers the threshold for rRNA-driven phase separation[17,22] and, in cells, exhibits higher partitioning with slower FRAP (stronger retention)[22]. In contrast, NMP1 mutB2 (and to a lesser extent NMP1 mutB1) reduces phase separation capacity in vitro[17] and, in cells, lowers partitioning with faster FRAP, severely for B2 and modestly for B1[22]. Therefore, mutations that make NMP1 more prone to phase separate in vitro show increased partitioning (or retention) of NMP1 in the nucleolus, and on the other hand, mutants that reduce the ability to phase separate in vitro show lower partitioning (or retention) in the nucleolus[17,22]. Extending this logic, one may hypothesize that age-driven shifts toward slower FRAP and nucleolar amyloid-like signatures may reflect increased network connectivity that alters partitioning rules, for instance: whether a protein enters, is retained within, or excluded from the nucleolus. In parallel, mesoscale compartmentalization is rewired: subphase boundaries blur and coarsen (fewer but larger bodies; a denser inner layer), eroding the layered architecture that normally orders rRNA processing and repair routing. In our work, expanded nucleoli in aged cells lose selectivity and nuclear partitioning, admitting entry to proteins normally excluded from small, young nucleoli (e.g., Rad52 and inert GFP), with functional costs for genome maintenance[9]. Consistent with this idea, Na-butyrate–induced senescent human cells revealed selective protein relocalization: NPM1 was depleted from nucleoli (accumulating in nucleoplasm), while PARP1 and DHX9 (non-nucleolar proteins) became enriched in nucleoli[80]. This phenotype was observed in the absence of shutdown of rRNA synthesis, hinting at changes in partitioning and
8. Concluding Remarks and Perspectives
Deciphering the mechanisms of aging has become a high priority in the context of a rapidly aging global population. To provide a conceptual structure for the expanding field of aging research, the “hallmarks of aging” framework was proposed, defining a dozen core processes that contribute to age-associated decline[1]. The nucleolus directly or indirectly intersects with most of these hallmarks through its central role in ribosome biogenesis and additional regulatory functions. Moreover, the repetitive nature and extremely high transcriptional activity of rDNA, necessary to meet cellular ribosomal demand, render it particularly vulnerable to genomic instability, itself a hallmark of aging[1]. The nucleolus also integrates signals from numerous well-studied pro-longevity interventions[29], modulating its architecture in response.
Recent studies have highlighted that the biophysical properties of the nucleolus, including its material state, internal compartmentalization, and the partitioning of the nuclear/nucleolar proteins, are critical for the precision and robustness of its functions. Although nucleolar size does not directly reflect the condensation state of the nucleolus, it is a readily quantifiable variable that often correlates with altered nucleolar properties during aging[8,9,16,26,73]. Increasing evidence suggests that aberrant phase transitions may contribute to, rather than merely accompany, the aging process[9,16,73].
It is obvious that the changes in the nucleolar condensate are driving rDNA instability and limit lifespan in budding yeast mitotic aging[9], whether nucleolar expansion acts as a driver of organismal aging in mammals remains unresolved and will require direct manipulation of nucleolar size or phase state. Dissecting the precise changes in nucleolar phase behavior during aging, and their effects across different cell types, is ongoing work that may reveal novel targets for pro-longevity interventions. One intriguing possibility is the development of drugs that directly target nucleolar phase behavior to “rejuvenate” the nucleolus in aged cells. Although therapeutics that modulate the condensation state of membraneless organelles remain in their infancy, the nucleolus stands out as a strong candidate for intervention.
Authors contribution
Gutierrez JI: Writing-original draft.
Tyler JK: Writing review & editing.
Conflicts of interest
Not applicable.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Availability of data and materials
Not applicable.
Funding
The work was funded by NIH (Grant No. RO1 AG079883).
Copyright
© The Author(s) 2026.
References
-
1. López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. Hallmarks of aging: An expanding universe. Cell. 2023;186(2):243-278.[DOI]
-
2. Fontana L, Partridge L. Promoting health and longevity through diet: From model organisms to humans. Cell. 2015;161(1):106-118.[DOI]
-
3. Lee H, Lee SV. Recent progress in regulation of aging by insulin/IGF-1 signaling in caenorhabditis elegans. Mol Cells. 2022;45(11):763-770.[DOI]
-
4. Zhang H, Xiao X, Pan Z, Dokudovskaya S. mTOR signaling networks: Mechanistic insights and translational frontiers in disease therapeutics. Signal Transduct Target Ther. 2025;10:428.[DOI]
-
5. Ripa R, Ballhysa E, Steiner JD, Laboy R, Annibal A, Hochhard N, et al. Refeeding-associated AMPKγ1 complex activity is a hallmark of health and longevity. Nat Aging. 2023;3(12):1544-1560.[DOI]
-
6. Ponti D. The nucleolus: A central hub for ribosome biogenesis and cellular regulatory signals. Int J Mol Sci. 2025;26(9):4174.[DOI]
-
7. Bemiller PM, Lee LH. Nucleolar changes in senescing WI-38 cells. Mech Ageing Dev. 1978;8:417-427.[DOI]
-
8. Tiku V, Jain C, Raz Y, Nakamura S, Heestand B, Liu W, et al. Small nucleoli are a cellular hallmark of longevity. Nat Commun. 2017;8:16083.[DOI]
-
9. Gutierrez JI, Tyler JK. A mortality timer based on nucleolar size triggers nucleolar integrity loss and catastrophic genomic instability. Nat Aging. 2024;4(12):1782-1793.[DOI]
-
10. Brangwynne CP, Mitchison TJ, Hyman AA. Active liquid-like behavior of nucleoli determines their size and shape in Xenopus laevis oocytes. Proc Natl Acad Sci U S A. 2011;108(11):4334-4339.[DOI]
-
11. Berry J, Weber SC, Vaidya N, Haataja M, Brangwynne CP. RNA transcription modulates phase transition-driven nuclear body assembly. Proc Natl Acad Sci U S A. 2015;112(38):E5237-E5245.[DOI]
-
12. Lafontaine DLJ, Riback JA, Bascetin R, Brangwynne CP. The nucleolus as a multiphase liquid condensate. Nat Rev Mol Cell Biol. 2021;22(3):165-182.[DOI]
-
13. Frottin F, Schueder F, Tiwary S, Gupta R, Körner R, Schlichthaerle T, et al. The nucleolus functions as a phase-separated protein quality control compartment. Science. 2019;365(6451):342-347.[DOI]
-
14. Montgomery Jr TSH. Comparative cytological studies, with especial regard to the morphology of the nucleolus. J Morphol. 1898;15(2):265-582.[DOI]
-
15. Smetana K, Daskal I, Busch H. Cytochemistry of the microtrabecular network in compact nucleoli of hepatocytes treated with cycloheximide. Histochemistry. 1980;65(3):301-308.[DOI]
-
16. Feric M, Vaidya N, Harmon TS, Mitrea DM, Zhu L, Richardson TM, et al. Coexisting liquid phases underlie nucleolar subcompartments. Cell. 2016;165(7):1686-1697.[DOI]
-
17. Mitrea DM, Cika JA, Stanley CB, Nourse A, Onuchic PL, Banerjee PR, et al. Self-interaction of NPM1 modulates multiple mechanisms of liquid–liquid phase separation. Nat Commun. 2018;9:842.[DOI]
-
18. Riback JA, Zhu L, Ferrolino MC, Tolbert M, Mitrea DM, Sanders DW, et al. Composition-dependent thermodynamics of intracellular phase separation. Nature. 2020;581(7807):209-214.[DOI]
-
19. Amin MA, Matsunaga S, Uchiyama S, Fukui K. Depletion of nucleophosmin leads to distortion of nucleolar and nuclear structures in HeLa cells. Biochem J. 2008;415(3):345-351.[DOI]
-
20. Koh CM, Gurel B, Sutcliffe S, Aryee MJ, Schultz D, Iwata T, et al. Alterations in nucleolar structure and gene expression programs in prostatic neoplasia are driven by the MYC oncogene. Am J Pathol. 2011;178(4):1824-1834.[DOI]
-
21. Martin RM, Ter-Avetisyan G, Herce HD, Ludwig AK, Lättig-Tünnemann G, Cardoso MC. Principles of protein targeting to the nucleolus. Nucleus. 2015;6(4):314-325.[DOI]
-
22. Sheu-Gruttadauria J, Yan X, Stuurman N, Vale RD, Floor SN. Nucleolar dynamics are determined by the ordered assembly of the ribosome. bioRxiv 559432 [Preprint]. 2024.[DOI]
-
23. King MR, Ruff KM, Lin AZ, Pant A, Farag M, Lalmansingh JM, et al. Macromolecular condensation organizes nucleolar sub-phases to set up a pH gradient. Cell. 2024;187(8):1889-1906.e24.[DOI]
-
24. Riback JA, Eeftens JM, Lee DSW, Quinodoz SA, Donlic A, Orlovsky N, et al. Viscoelasticity and advective flow of RNA underlies nucleolar form and function. Mol Cell. 2023;83(17):3095-3107.[DOI]
-
25. Bemiller PM, Miller JE. Cytological changes in senescing WI-38 cells: A statistical analysis. Mech Ageing Dev. 1979;10:1-15.[DOI]
-
26. Buchwalter A, Hetzer MW. Nucleolar expansion and elevated protein translation in premature aging. Nat Commun. 2017;8:328.[DOI]
-
27. Sinclair DA, Mills K, Guarente L. Accelerated aging and nucleolar fragmentation in Yeast sgs1 Mutants. Science. 1997;277(5330):1313-1316.[DOI]
-
28. Sirozh O, Saez-Mas A, Jung B, Sanchez-Burgos L, Zarzuela E, Rodrigo-Perez S, et al. Nucleolar stress caused by arginine-rich peptides triggers a ribosomopathy and accelerates aging in mice. Mol Cell. 2024;84(8):1527-1540.[DOI]
-
29. Tiku V, Antebi A. Nucleolar function in lifespan regulation. Trends Cell Biol. 2018;28(8):662-672.[DOI]
-
30. Ünal E, Kinde B, Amon A. Gametogenesis eliminates age-induced cellular damage and resets life span in yeast. Science. 2011;332(6037):1554-1557.[DOI]
-
31. Kodan N, Hussaini R, Weber SC, Kondev J, Mohapatra L. Transcription-templated assembly of the nucleolus in the Caenorhabditis elegans embryo. Proc Natl Acad Sci U S A. 2025;122(22):e2411964122.[DOI]
-
32. Quinodoz SA, Jiang L, Abu-Alfa AA, Comi TJ, Zhao H, Yu Q, et al. Mapping and engineering RNA-driven architecture of the multiphase nucleolus. Nature. 2025;644(8076):557-566.[DOI]
-
33. Morlot S, Song J, Léger-Silvestre I, Matifas A, Gadal O, Charvin G. Excessive rDNA transcription drives the disruption in nuclear homeostasis during entry into senescence in budding yeast. Cell Rep. 2019;28(2):408-422.[DOI]
-
34. Ubaida-Mohien C, Lyashkov A, Gonzalez-Freire M, Tharakan R, Shardell M, Moaddel R, et al. Discovery proteomics in aging human skeletal muscle finds change in spliceosome, immunity, proteostasis and mitochondria. eLife. 2019;8:e49874.[DOI]
-
35. Ward W, Richardson A. Effect of age on liver protein synthesis and degradation. Hepatology. 1991;14(5):935-948.[DOI]
-
36. Kelmer Sacramento E, Kirkpatrick JM, Mazzetto M, Baumgart M, Bartolome A, Di Sanzo S, et al. Reduced proteasome activity in the aging brain results in ribosome stoichiometry loss and aggregation. Mol Syst Biol. 2020;16(6):e9596.[DOI]
-
37. Tapia O, Narcís JO, Riancho J, Tarabal O, Piedrafita L, Calderó J, et al. Cellular bases of the RNA metabolism dysfunction in motor neurons of a murine model of spinal muscular atrophy: Role of Cajal bodies and the nucleolus. Neurobiol Dis. 2017;108:83-99.[DOI]
-
38. Kelmer Sacramento E, Kirkpatrick JM, Mazzetto M, Baumgart M, Bartolome A, Di Sanzo S, et al. Reduced proteasome activity in the aging brain results in ribosome stoichiometry loss and aggregation. Mol Syst Biol. 2020;16(6):e9596.[DOI]
-
39. Ali A, Garde R, Schaffer OC, Bard JAM, Husain K, Kik SK, et al. Adaptive preservation of orphan ribosomal proteins in chaperone-dispersed condensates. Nat Cell Biol. 2023;25(11):1691-1703.[DOI]
-
40. Dogra P, Ferrolino MC, Khatun S, Tolbert M, Miao Q, Pruett-Miller SM, et al. Granular component sub-phases direct ribosome biogenesis in the nucleolus. bioRxiv 640913 [Preprint]. 2025.[DOI]
-
41. Okuwaki M, Saito S, Hirawake-Mogi H, Nagata K. The interaction between nucleophosmin/NPM1 and the large ribosomal subunit precursors contribute to maintaining the nucleolar structure. Biochim Biophys Acta Mol Cell Res. 2021;1868(1):118879.[DOI]
-
42. Akamatsu Y, Kobayashi T. The human RNA polymerase I transcription terminator complex acts as a replication fork barrier that coordinates the progress of replication with rRNA transcription activity. Mol Cell Biol. 2015;35(10):1871-1881.[DOI]
-
43. Sinclair DA, Guarente L. Extrachromosomal rDNA circles—A cause of aging in yeast. Cell. 1997;91(7):1033-1042.[DOI]
-
44. Pal S, Postnikoff SD, Chavez M, Tyler JK. Impaired cohesion and homologous recombination during replicative aging in budding yeast. Sci Adv. 2018;4(2):eaaq0236.[DOI]
-
45. Nelson JO, Slicko A, Yamashita YM. The retrotransposon R2 maintains Drosophila ribosomal DNA repeats. Proc Natl Acad Sci U S A. 2023;120(23):e2221613120.[DOI]
-
46. Lu KL, Nelson JO, Watase GJ, Warsinger-Pepe N, Yamashita YM. Transgenerational dynamics of rDNA copy number in Drosophila male germline stem cells. eLife. 2018;7:e32421.[DOI]
-
47. Watada E, Li S, Hori Y, Fujiki K, Shirahige K, Inada T, et al. Age-dependent ribosomal DNA variations in mice. Mol Cell Biol. 2020;40(22):e00368-e00320.[DOI]
-
48. Ren R, Deng L, Xue Y, Suzuki K, Zhang W, Yu Y, et al. Visualization of aging-associated chromatin alterations with an engineered TALE system. Cell Res. 2017;27(4):483-504.[DOI]
-
49. Hallgren J, Pietrzak M, Rempala G, Nelson PT, Hetman M. Neurodegeneration-associated instability of ribosomal DNA. Biochim Biophys Acta Mol Basis Dis. 2014;1842(6):860-868.[DOI]
-
50. Hori Y, Shimamoto A, Kobayashi T. The human ribosomal DNA array is composed of highly homogenized tandem clusters. Genome Res. 2021;31(11):1971-1982.[DOI]
-
51. Hu Z, Chen K, Xia Z, Chavez M, Pal S, Seol JH, et al. Nucleosome loss leads to global transcriptional up-regulation and genomic instability during yeast aging. Genes Dev. 2014;28(4):396-408.[DOI]
-
52. Møller HD, Mohiyuddin M, Prada-Luengo I, Sailani MR, Halling JF, Plomgaard P, et al. Circular DNA elements of chromosomal origin are common in healthy human somatic tissue. Nat Commun. 2018;9:1069.[DOI]
-
53. Cohen S, Agmon N, Sobol O, Segal D. Extrachromosomal circles of satellite repeats and 5S ribosomal DNA in human cells. Mob DNA. 2010;1(1):11.[DOI]
-
54. Tchurikov NA, Yudkin DV, Gorbacheva MA, Kulemzina AI, Grischenko IV, Fedoseeva DM, et al. Hot spots of DNA double-strand breaks in human rDNA units are produced in vivo. Sci Rep. 2016;6:25866.[DOI]
-
55. Paredes S, Angulo-Ibanez M, Tasselli L, Carlson SM, Zheng W, Li TM, et al. The epigenetic regulator SIRT7 guards against mammalian cellular senescence induced by ribosomal DNA instability. J Biol Chem. 2018;293(28):11242-11250.[DOI]
-
56. Torres-Rosell J, Sunjevaric I, De Piccoli G, Sacher M, Eckert-Boulet N, Reid R, et al. The Smc5–Smc6 complex and SUMO modification of Rad52 regulates recombinational repair at the ribosomal gene locus. Nat Cell Biol. 2007;9(8):923-931.[DOI]
-
57. van Sluis M, McStay B. A localized nucleolar DNA damage response facilitates recruitment of the homology-directed repair machinery independent of cell cycle stage. Genes Dev. 2015;29(11):1151-1163.[DOI]
-
58. Welty S, Teng Y, Liang Z, Zhao W, Sanders LH, Greenamyre JT, et al. RAD52 is required for RNA-templated recombination repair in
post-mitotic neurons. J Biol Chem. 2018;293(4):1353-1362.[DOI] -
59. Alberti S, Gladfelter A, Mittag T. Considerations and challenges in studying liquid-liquid phase separation and biomolecular condensates. Cell. 2019;176(3):419-434.[DOI]
-
60. Banani SF, Lee HO, Hyman AA, Rosen MK. Biomolecular condensates: Organizers of cellular biochemistry. Nat Rev Mol Cell Biol. 2017;18(5):285-298.[DOI]
-
61. Noel JS, Dewey WC, Abel JH Jr, Thompson RP. Ultrastructure of the nucleolus during the Chinese hamster cell cycle. J Cell Biol. 1971;49(3):830-847.[DOI]
-
62. Abraham KJ, Khosraviani N, Chan JNY, Gorthi A, Samman A, Zhao DY, et al. Nucleolar RNA polymerase II drives ribosome biogenesis. Nature. 2020;585(7824):298-302.[DOI]
-
63. Audas TE, Audas DE, Jacob MD, Ho JJD, Khacho M, Wang M, et al. Adaptation to stressors by systemic protein amyloidogenesis. Dev Cell. 2016;39(2):155-168.[DOI]
-
64. Audas TE, Jacob MD, Lee S. Immobilization of proteins in the nucleolus by ribosomal intergenic spacer noncoding RNA. Mol Cell. 2012;45(2):147-157.[DOI]
-
65. Alberti S, Hyman AA. Are aberrant phase transitions a driver of cellular aging? BioEssays. 2016;38(10):959-968.[DOI]
-
66. Düster R, Kaltheuner IH, Schmitz M, Geyer M. 1,6-Hexanediol, commonly used to dissolve liquid–liquid phase separated condensates, directly impairs kinase and phosphatase activities. J Biol Chem. 2021;296:100260.[DOI]
-
67. Kroschwald S, Maharana S, Mateju D, Malinovska L, Nüske E, Poser I, et al. Promiscuous interactions and protein disaggregases determine the material state of stress-inducible RNP granules. eLife. 2015;4:e06807.[DOI]
-
68. Hayflick L, Moorhead PS. The serial cultivation of human diploid cell strains. Exp Cell Res. 1961;25(3):585-621.[DOI]
-
69. López-Gil L, Pascual-Ahuir A, Proft M. Genomic instability and epigenetic changes during aging. Int J Mol Sci. 2023;24(18):14279.[DOI]
-
70. Hernandez-Segura A, Nehme J, Demaria M. Hallmarks of cellular senescence. Trends Cell Biol. 2018;28(6):436-453.[DOI]
-
71. Yang L, Song T, Chen L, Soliman H, Chen J. Nucleolar repression facilitates initiation and maintenance of senescence. Cell Cycle. 2015;14(22):3613-3623.[DOI]
-
72. Chen J, Lobb IT, Morin P, Novo SM, Simpson J, Kennerknecht K, et al. Identification of a novel TIF-IA–NF-κB nucleolar stress response pathway. Nucleic Acids Res. 2018;46(12):6188-6205.[DOI]
-
73. Jo M, Kim S, Park J, Chang YT, Gwon Y. Reduced dynamicity and increased high-order protein assemblies in dense fibrillar component of the nucleolus under cellular senescence. Redox Biol. 2024;75:103279.[DOI]
-
74. Ishibashi S, Inoko A, Oka Y, Leproux P, Kano H. Coherent Raman microscopy visualizes ongoing cellular senescence through amide I peak shifts originating from β sheets in disordered nucleolar proteins. Sci Rep. 2024;14:27584.[DOI]
-
75. Soto C, Pritzkow S. Protein misfolding, aggregation, and conformational strains in neurodegenerative diseases. Nat Neurosci. 2018;21(10):1332-1340.[DOI]
-
76. Lourenco GF, Torres-Pacheco ME, Fu Y, Li H, McCann H, Shepherd CE, et al. Nucleolar aggregation of key neuropathological proteins in the postmortem neurodegenerative brain. Acta Neuropathol. 2025;150:60.[DOI]
-
77. Yu H, Sun Z, Tan T, Pan H, Zhao J, Zhang L, et al. rRNA biogenesis regulates mouse 2C-like state by 3D structure reorganization of
peri-nucleolar heterochromatin. Nat Commun. 2021;12:6365.[DOI] -
78. Hyman AA, Weber CA, Jülicher F. Liquid-liquid phase separation in biology. Annu Rev Cell Dev Biol. 2014;30:39-58.[DOI]
-
79. Ferrolino MC, Mitrea DM, Michael JR, Kriwacki RW. Compositional adaptability in NPM1-SURF6 scaffolding networks enabled by dynamic switching of phase separation mechanisms. Nat Commun. 2018;9:5064.[DOI]
-
80. Kar B, Liu B, Zhou Z, Lam YW. Quantitative nucleolar proteomics reveals nuclear re-organization during stress-induced senescence in mouse fibroblast. BMC Cell Biol. 2011;12:33.[DOI]
-
81. Cochard A, Garcia-Jove Navarro M, Piroska L, Kashida S, Kress M, Weil D, et al. RNA at the surface of phase-separated condensates impacts their size and number. Biophys J. 2022;121(9):1675-1690.[DOI]
-
82. Ma W, Zhen G, Xie W, Mayr C. In vivo reconstitution finds multivalent RNA–RNA interactions as drivers of mesh-like condensates. eLife. 2021;10:e64252.[DOI]
-
83. Rana U, Xu K, Narayanan A, Walls MT, Panagiotopoulos AZ, Avalos JL, et al. Asymmetric oligomerization state and sequence patterning can tune multiphase condensate miscibility. Nat Chem. 2024;16(7):1073-1082.[DOI]
-
84. Farag M, Cohen SR, Borcherds WM, Bremer A, Mittag T, Pappu RV. Condensates formed by prion-like low-complexity domains have small-world network structures and interfaces defined by expanded conformations. Nat Commun. 2022;13:7722.[DOI]
-
85. Mittag T, Pappu RV. A conceptual framework for understanding phase separation and addressing open questions and challenges. Mol Cell. 2022;82(12):2201-2214.[DOI]
-
86. Kelley FM, Ani A, Pinlac EG, Linders B, Favetta B, Barai M, et al. Controlled and orthogonal partitioning of large particles into biomolecular condensates. Nat Commun. 2025;16:3521.[DOI]
-
87. Dar F, Cohen SR, Mitrea DM, Phillips AH, Nagy G, Leite WC, et al. Biomolecular condensates form spatially inhomogeneous network fluids. Nat Commun. 2024;15:3413.[DOI]
Copyright
© The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Publisher’s Note
Share And Cite



