Haiyan Yan, School of Medicine, Hangzhou City University, Hangzhou 310015, Zhejiang, China. E-mail: yanhy@hzcu.edu.cn
Abstract
Sister chromatid cohesion, established during DNA replication, is essential for accurate chromosome segregation. While acetylation of the cohesin subunit SMC3 by ESCO1/2 promotes the recruitment of the cohesin stabilizer Sororin, this pathway is insufficient for full Sororin function. In a recent study, Jiang et al. identify a previously uncharacterized human microprotein, RSMC, as a Sororin cofactor required for sister chromatid cohesion. The authors show that RSMC interacts with Sororin, and this interaction is enhanced during S-phase by PARP1/2-mediated poly(ADP-ribosyl)ation (PARylation) of RSMC. PARylation of RSMC, triggered by DNA replication intermediates, acts in parallel with SMC3 acetylation to ensure the timely and efficient recruitment of Sororin to chromatin. Consequently, inhibition of PARP activity reduces chromatin-bound Sororin and causes cohesion defects, which can be rescued by overexpressing wild-type RSMC or Sororin, but not by PARylation- or interaction-deficient mutants. Furthermore, RSMC stimulates the anti-Wapl activity of Sororin in vitro, promoting stable cohesin binding. Taken together, the work of Jiang et al. describes a dual, replication-coupled regulatory mechanism wherein S-phase PARylation of the microprotein RSMC cooperates with SMC3 acetylation to fully enable Sororin’s function in establishing sister chromatid cohesion. This mechanism is important for maintaining genomic stability, and its dysregulation may contribute to chromosome segregation errors observed in cancer.
Keywords
The faithful segregation of genetic material during mitosis is a cornerstone of genomic stability and is critically dependent on the establishment of sister chromatid cohesion during S-phase[1]. This process is tightly coupled to DNA replication, with the replication fork serving as a central signaling hub where multiple pathways converge to ensure timely and accurate cohesion. While the cohesin complex and its regulator Sororin, which acts downstream of SMC3 acetylation, are well-characterized, the full complement of signals ensuring Sororin’s efficient, replication-coupled recruitment has remained incomplete[2-4]. The study by Jiang et al. addresses this gap by identifying a parallel regulatory axis involving the DNA damage-associated enzyme PARP1 and a novel microprotein cofactor, RSMC (Figure 1)[5]. Notably, the PARP activity described here is distinct from its canonical role in the DNA damage response; rather, it is triggered by unligated Okazaki fragments and other replication intermediates during normal S-phase, representing a replication-coupled signaling function that directly regulates chromosome architecture. This work reveals a role for PARP activity in unperturbed S-phase progression and highlights the functional potential of microproteins, an emerging class of polypeptides derived from alternative open reading frames[6]. Given the clinical use of PARP inhibitors and the recognition of cohesinopathies in diseases such as Cornelia de Lange syndrome (CdLS; the most prevalent), Roberts syndrome (RBS), and Warsaw breakage syndrome (WABS), this research offers mechanistic insights with potential implications.
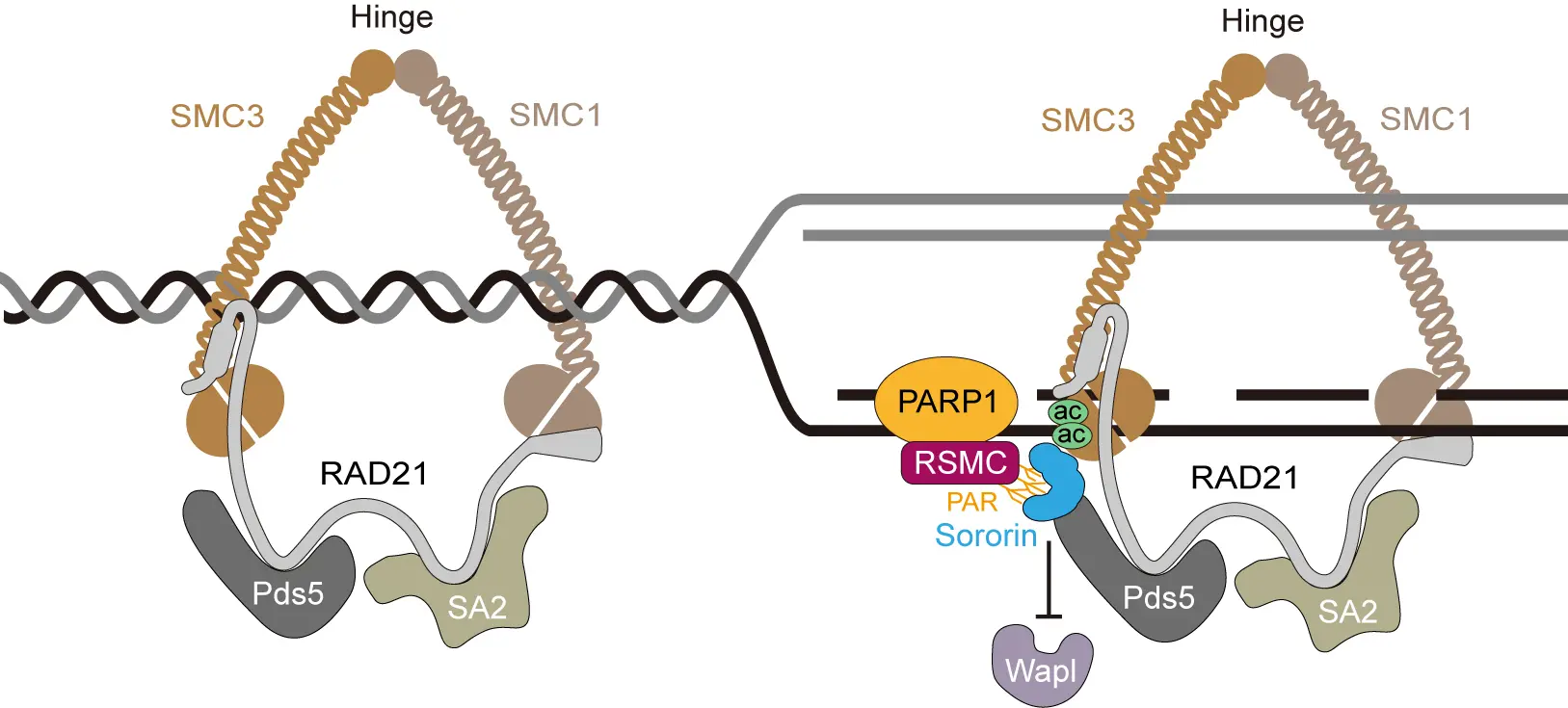
Figure 1. A model showing how PARylation at the fork allows RSMC to bridge replication to Sororin-mediated sister chromatid cohesion. SMC1: structural maintenance of chromosomes 1; SMC3: structural maintenance of chromosomes 3; RAD21: radiation sensitive 21; SA2: stromal antigen 2; Pds5: precocious dissociation of sisters 5; Wapl: wings apart-like protein; PARP1: poly(ADP-ribose) polymerase 1; PAR: poly(ADP-ribose).
The authors begin by succinctly reviewing the cohesin complex and the established pathway wherein ESCO1/2-mediated SMC3 acetylation promotes Sororin recruitment[2,7]. They note the evidence from prior literature that SMC3 acetylation is necessary but insufficient for robust Sororin function, indicating that regulators are missing[2]. This premise provides a logical rationale for a discovery-based approach to identify novel Sororin-interacting factors.
The experimental narrative follows a logical progression. The identification of RSMC via immunoprecipitation-mass spectrometry (IP-MS) is a standard starting point, validated through reciprocal co-immunoprecipitation and in situ proximity ligation assays (PLA). The functional requirement for RSMC is demonstrated through knockdown, heterozygous knockout, complementation, and both classical (chromosome spreads) and modern (CRISPR/dCas9-based) phenotyping assays. The structural and functional dissection of the RSMC-Sororin interface is thorough. Guided by AlphaFold2 and GRAMM predictions, the authors map interaction motifs and generate defective mutants (RSMCΔSIM, RSMC-16A). The failure of these mutants to rescue cohesion defects indicates the functional importance of the interaction.
The regulation of this interaction by PARP1-mediated PARylation is uncovered through a series of logical experiments. The demonstration of a cell-cycle-regulated, PARP-activity-dependent interaction between PARP1 and RSMC, followed by in vitro and in vivo PARylation assays, is systematic. The coincidence of PARylation sites with the Sororin-interacting motifs links the modification to function. Biochemical assays showing enhanced Sororin binding to PARylated RSMC further support the proposed mechanism.
The authors integrate the new PARP-RSMC axis with the established SMC3ac pathway. Data showing additive effects on chromatin-bound Sororin loss and cohesion defects upon combined perturbation of RSMC and ESCO1/2 support the model of two independent, cooperative pathways. The finding that overexpression of wild-type RSMC or Sororin, but not PAR-binding or PARylation-deficient mutants, rescues cohesion defects induced by PARP inhibition identifies RSMC as the relevant PARP target. The demonstration via microscale thermophoresis (MST) that RSMC stimulates Sororin’s anti-Wapl activity provides a plausible biochemical explanation for the cellular phenotype.
The authors synthesize the findings into a coherent “dual-pathway” model and contextualize them within the literature on PARP function, Okazaki fragment processing, synthetic lethality, and microprotein biology[8-10]. They also appropriately acknowledge the separate role of RSMC in ribosome biogenesis[11], noting an intriguing question for future work.
A key unresolved question is the precise structural mechanism by which PARylation enhances the RSMC-Sororin-PDS5 interaction. While the functional consequences of RSMC PARylation are convincingly demonstrated, the atomic-level details of how PAR chains modulate the protein–protein interface remain unknown. Future structural studies, for example, solving the architecture of the PARylated RSMC-Sororin complex, will be essential to understand this regulation. Beyond this, some potential caveats could be further explored. While the concordance between PARP inhibition and genetic depletion is strong, the individual contributions of PARP1 versus PARP2 in vivo could be explored further. The precise structural mechanism by which PARylation enhances the RSMC-Sororin-PDS5 interaction remains unresolved. The authors could also briefly consider whether RSMC PARylation affects its own stability or localization independently of Sororin binding.
This work advances understanding in several areas. First, it expands the regulatory landscape of sister chromatid cohesion by establishing PARylation as a parallel, replication-coupled signal alongside acetylation. This extends the known functional repertoire of PARP enzymes beyond DNA damage responses to include direct regulation of chromosome architecture during normal S-phase. Second, it illustrates a mechanism for signal integration. The requirement for two independent, replication-coupled modifications to fully activate Sororin could ensure fidelity by coupling cohesion establishment strictly to ongoing DNA synthesis. Third, it demonstrates the functional significance of a microprotein. RSMC serves as an example of how small, alternative ORF-encoded polypeptides can occupy specific niches within core cellular machineries.
The findings of Jiang et al. are relevant to contexts like aging and cancer, where genomic instability is a hallmark and defective DNA replication and chromosome segregation are prevalent. The identified PARP1-RSMC-Sororin axis presents a molecular link between replication fork signaling, cohesion establishment, and genomic integrity. In aging, a decline in DNA repair efficiency and accumulation of replication stress have been documented. A compromised PARP-RSMC pathway could contribute to sub-optimal sister chromatid cohesion, potentially leading to increased chromosome mis-segregation, aneuploidy, and age-related functional decline. This study invites investigation into whether this pathway deteriorates with age.
For cancer biology, the implications are twofold. Cohesion factors are frequently mutated, and the discovery of RSMC adds a new potential player whose dysregulation could promote chromosomal instability. Furthermore, the work provides a perspective on the mechanism of PARP inhibitors (PARPi). While clinical efficacy is primarily attributed to synthetic lethality, this study reveals that PARPi also disrupt a replication-coupled process required for accurate mitosis. This effect on cohesion could contribute to the drugs’ cytostatic and cytotoxic effects. Specifically, PARPi-induced impairment of the RSMC-Sororin interaction reduces chromatin-bound Sororin, leading to defective sister chromatid cohesion. This can result in (i) mitotic errors such as premature chromatid separation and chromosome mis-segregation, which promote aneuploidy and genomic instability; (ii) resulting cohesion failure can trigger prolonged mitotic arrest, mitotic catastrophe, or apoptosis, underpinning the cytostatic and cytotoxic effects of PARPi; (iii) moreover, since the PARP-RSMC-Sororin axis operates in normal S-phase progression, its inhibition in healthy proliferating tissues may contribute to treatment-associated side effects, such as bone marrow suppression, by compromising cohesion and chromosome segregation in normal cells. Understanding the RSMC pathway may help explain variable patient responses to PARPi and inform strategies to manage therapeutic resistance or toxicity.
In cohesin biology, this study will likely prompt re-examination of phenotypes associated with PARP inhibition to include potential cohesion deficiencies. It may inspire searches for similar microprotein cofactors in other aspects of cohesin dynamics. For PARP biology, the paper underscores the importance of basal S-phase PARP activity, suggesting that the efficacy and toxicity of PARP inhibitors may stem partly from disruption of fundamental processes like cohesion. Finally, it provides a rationale for further characterization of microproteins, with potential implications for understanding physiology and disease.
This work also opens numerous avenues for future research. Mechanistically, solving the structure of the PARylated RSMC-Sororin complex is a clear next step. The spatial regulation of this pathway at replication forks and its potential coordination with other fork-associated events remain to be explored. The link between RSMC’s roles in ribosome biogenesis and cohesion raises the possibility that this microprotein could coordinate biosynthetic capacity with cell cycle progression.
While the evidence is strong, some caution is warranted. The reliance on RSMC haploinsufficiency suggests potential embryonic lethality for the full knockout, implying essential functions beyond cohesion that could influence phenotypic interpretations. The rescue by RSMC overexpression does not entirely exclude contributions from other PARP substrates in the cohesion process. Furthermore, the physiological stoichiometry and temporal dynamics of the RSMC-Sororin complex in vivo warrant further investigation.
In conclusion, Jiang et al. have identified a missing regulator in the sister chromatid cohesion pathway, revealing a PARP-dependent axis that operates in parallel to the canonical acetylation pathway. By centering this discovery on a microprotein, they have both addressed a specific biological question and provided an example of the functional potential within the unexplored proteome. This work deepens understanding of chromosome dynamics, expands the known physiological roles of PARPs, and will likely stimulate cross-disciplinary research at the intersection of cell cycle regulation, chromatin biology, and genomics. Its contribution is a step toward a more integrated understanding of how fundamental cellular processes are regulated through multi-layered control and how their dysregulation may contribute to human disease.
Acknowledgments
We thank members of our lab for helpful discussions. ChatGPT was used solely for language polishing. The authors take full responsibility for the final manuscript.
Authors contribution
Yuan J, Chen Q, Wang F, Yan H: Writing-original draft.
Conflicts of interest
The authors declared that there are no conflicts of interest.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Availability of data and materials
Not applicable.
Funding
This work was supported by grants from the National Natural Science Foundation of China (32270772, W2511031, 32025011, and 32100583), the Natural Science Foundation of Zhejiang Province (LZ24C070001 and MS26C070001), and the National Key Research & Development Program of China (2022YFA1105200 and 2022YFA1303100).
Copyright
© The Author(s) 2026.
References
-
1. Nasmyth K, Haering CH. The structure and function of smc and kleisin complexes. Annu Rev Biochem. 2005;74:595-648.[DOI]
-
5. Jiang M, Zhang J, He J, Miao Y, Wang L, Zhong H, et al. S-phase PARylation of microprotein RSMC enhances the function of Sororin in sister chromatid cohesion. EMBO J. 2026;45(1):278-309.[DOI]
Copyright
© The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Publisher’s Note
Share And Cite




