Bo Ying, Suzhou Abogen Biosciences Inc. Ltd., Suzhou 215000, Jiangsu, China. E-mail: bo.ying@abogenbio.com
Abstract
Artificial intelligence (AI) is revolutionizing the design of ionizable lipids, the pivotal components of lipid nanoparticles (LNPs) for messenger RNA (mRNA) delivery, enabling efficient exploration of vast chemical space of ionizable lipids beyond the reach of traditional methods. This mini-review explores the burgeoning field of AI-powered design and optimization of ionizable lipids for mRNA delivery. We also discuss the critical role of high-throughput experimental strategies, particularly barcoding coupled with next-generation sequencing, in generating the large-scale in vivo datasets for model training. Finally, we discuss current challenges, including data quality and the necessity for domain-specific modeling strategies, and present a future outlook on the integration of AI with scientific computing for LNP research.
Keywords
1. Introduction
The remarkable success of messenger RNA (mRNA) vaccines during the COVID-19 pandemic has significantly advanced interest in mRNA-based vaccines and therapies, underscoring the critical role of their delivery platform, lipid nanoparticles (LNPs). Naked mRNA is inherently unstable and susceptible to degradation by serum ribonucleases, preventing it from reaching target cells[1]. LNPs are engineered to circumvent these barriers by encapsulating mRNA, facilitating its transport across extracellular matrices and cell membranes, and ultimately enabling its release into the cytosol[2,3].
A typical LNP formulation comprises four components[2,4]: ionizable lipids, helper lipids (phospholipids), cholesterol, and polyethylene glycol-lipids (PEG-lipids). Among these, ionizable lipids are paramount for delivery efficiency and potency. These lipids possess a unique pH-dependent charge characteristic: they are positively charged at acidic pH to complex with negatively charged mRNA, neutral at physiological pH to minimize toxicity during systemic circulation, and reprotonated in the acidic environment of late endosomes to promote endosomal membrane disruption and mRNA release[5,6].
Structurally, ionizable lipids consist of three modular parts[4]: a headgroup, a linker, and tail chains. The headgroup, typically containing ionizable amines (e.g., tertiary, primary, secondary amines, heterocycles, or guanidine), dictates the acid dissociation constant (pKa) of the lipid and influences the apparent pKa of the LNP[7]. This pKa is crucial for endosomal escape, with an optimal range (e.g., 6.2-6.6) being essential for high potency[1]. The headgroup also influences the toxicity profile. The linker connects the headgroup to the tails and primarily affects the biodegradability and metabolic stability of the lipid[2]. While various linkers exist (ethers, amides, phosphates), ester linkages are widely favored due to their facile biodegradation in vivo, leading to improved safety profiles[8]. The tail chains influence key properties such as pKa, membrane fluidity, fusogenicity, and overall potency[2,4]. Tail design involves variations in carbon chain length (typically 8-20 carbons), degree of unsaturation, symmetry, and branching. Aliphatic chains are common, but aromatic moieties like cholesterol derivatives can also be incorporated.
The collective geometry of these three components determines the lipid’s packing parameter (P)[2,9], defined as P = V/(a₀ × l), where V is the lipid tail volume, a₀ is the headgroup area, and l is the critical tail length. Lipids with a packing parameter P > 1, which adopt cone-shaped geometries, are more likely to form inverted micelles or hexagonal (HII) phases, facilitating endosomal escape and demonstrating superior mRNA delivery efficacy[10].
Given the profound impact of ionizable lipid structure on biodegradability, LNP pKa, transfection potency, and safety, there is a compelling need for their continuous optimization to develop superior mRNA delivery systems[7]. Furthermore, as mRNA therapeutics expand to target diverse tissues and diseases[11], the demand for novel ionizable lipids with tailored properties is growing. For instance, screening 56 headgroups on a constant dilinoleyl tail revealed that the headgroup significantly modulates LNP pKa and potency[12]. Similarly, varying tail lengths while keeping the headgroup constant can dramatically shift organ selectivity between the liver and spleen[13].
The evolution of ionizable lipids can be broadly categorized into three generations[1], reflecting a gradual shift from potency-driven design toward improved biodegradability, safety, and tissue-specific delivery. The first generation, exemplified by DLin-MC3-DMA (MC3) in Onpattro®, was developed through rational design and demonstrated high siRNA delivery efficacy[12], albeit with a long half-life (~72 hours)[1]. The second generation focused on enhancing biodegradability by incorporating ester linkages, leading to lipids like SM-102 (Moderna’s mRNA-1273)[14], ALC-0315 (Pfizer-BioNTech’s BNT162b2), and ATX-0126 (Arcturus’s ARCT-154). Current third-generation efforts aim to streamline synthesis, improve tissue-specific targeting, and enhance adaptability for various therapeutic modalities. Some representative ionizable lipids are listed in Table 1.
| Product Name | Sponsor | Ionizable Lipid | Structure | Ref. |
| Onpattro | Alnylam | DLin-MC3-DMA | 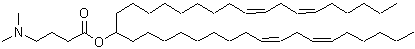 | [8] |
| BNT162b2 | Pfizer/BioNTech | ALC-0315 | 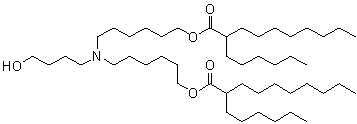 | [15] |
| mRNA-1273 | Moderna | SM-102 | 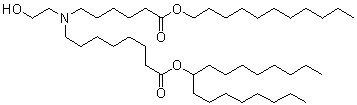 | [16] |
| ARCT-154 | Arcturus | ATX-0126 | 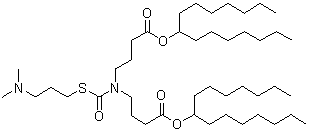 | [17] |
Ionizable lipid design strategies[18] include (1) rational design, which is effective but low-throughput and labor-intensive (e.g., MC3 development); (2) combinatorial chemistry, a high-throughput approach that addresses the throughput limitations of rational design but may lack structural novelty and relies heavily on empirical knowledge; and (3) in silico design[19], an emerging, powerful paradigm leveraging artificial intelligence (AI) for accelerated discovery, though it demands large, high-quality datasets, which can be a limiting factor.
This review focuses on the application of AI in optimizing ionizable lipids. We first introduce foundational computational methods in drug discovery. Second, we summarize current progress in AI-aided ionizable lipid design. Third, we discuss strategies for experimental data accumulation to fuel AI models. Finally, we address the current limitations and future perspectives in this burgeoning field.
2. Current Computational Methods in Drug Discovery
This section introduces cutting-edge computational methods in drug discovery, which form the foundation for the AI-aided design of ionizable lipids. An overview of the AI-assisted design and screening workflow for ionizable lipids is illustrated in Figure 1.
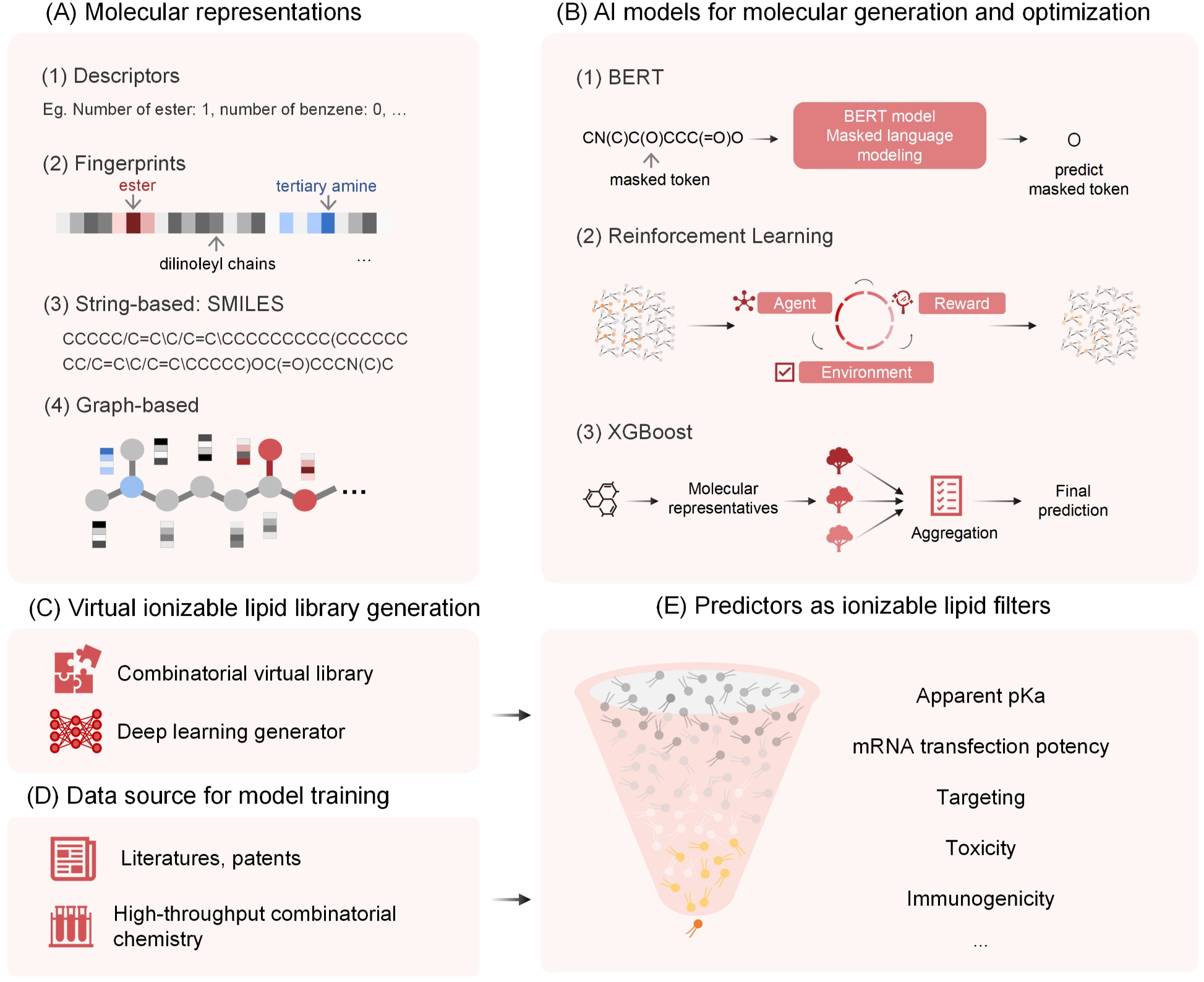
Figure 1. AI-assisted design and screening framework for ionizable lipids. AI: artificial intelligence; SMILES: simplified molecular input line entry system; XGBoost: extreme gradient boosting; BERT: bidirectional encoder representations from transformers; mRNA: messenger RNA.
2.1 Molecular representations for small molecules
Molecular representations are designed to translate structural information into a machine-readable format. The primary representations include descriptors, fingerprints, string-based notations, and graph-based representations. (1) Descriptors represent molecules using numerical values derived from their physicochemical properties. Common descriptors can be automatically calculated using toolkits such as RDKit, OpenBabel[20], and PaDEL[21]. (2) Fingerprints encode the presence or absence of specific chemical substructures within a molecule. Like descriptors, they can be readily generated using RDKit, OpenBabel, or PaDEL. Widely used fingerprints include extended-connectivity fingerprints (ECFP)[22], AtomPairs[23], and molecular access system (MACCS)[24] keys. (3) String-based representations, such as the simplified molecular input line entry system (SMILES)[25] and the international chemical identifier (InChI)[26], represent molecules as text strings. (4) Graph-based representations depict molecules as graphs, where atoms are represented as nodes and chemical bonds as edges[19,27]. This format naturally captures the molecular connectivity and topology.
2.2 Generative models
Generative models enable de novo drug design, moving beyond traditional, low-throughput rational design that relies heavily on expert experience[28]. This subsection introduces string-based and graph-based generative models, as well as reinforcement learning for molecular optimization. (1) String-based generative models treat molecular generation as a sequence-generation task, trained on SMILES strings to produce new, valid SMILES sequences[19]. Being data-driven, they can be pre-trained to generate molecules that are structurally similar to those in the training dataset. (2) Graph-based molecular generation. In contrast to sequence-based models, graph neural networks (GNNs) inherently encode molecular structure and positional information without requiring additional encoding schemes to represent graph connectivity[29], making them a natural fit for generating molecular graphs directly. (3) Reinforcement learning (RL) is typically employed for goal-directed optimization rather than de novo generation[30,31]. An RL agent iteratively proposes structural modifications and learns from the feedback provided by a reward function. While a generative model is trained to produce valid molecules, RL can be applied to optimize these molecules towards specific desirable properties, which are defined by the unique objectives of a given project.
3. AI-aided Ionizable Lipid Design
In the preceding section, we outlined recent advances in AI-aided drug discovery. Although AI has demonstrated considerable capability in small-molecule drug design, ionizable lipids present fundamentally different design challenges, as their biological performance emerges from collective nanoparticle-level properties rather than direct molecular target engagement. This is due to the need to predict key properties particularly relevant to LNPs, such as apparent pKa, mRNA transfection potency, and immunogenicity, which differ significantly from traditional small-molecule properties. Below we summarize notable studies employing computational and AI methods to facilitate ionizable lipid design (as partially illustrated in Table 2).
| Author | Dataset | Properties | Representation | Model Type | Task Type | Performance Metric | Year | Ref. |
| Dinesh M. Dhumal et al. | 50 (in vivo; Literature-curated) | Apparent pKa | Descriptors | PLS | Regressor | 93.3% correlation coefficient (test set) | 2020 | [32] |
| Wei Wang et al. | 325 (in vivo; Literature-curated) | IgG titer | ECFP | LightGBM | Regressor | R2 > 0.87 (10-fold cross validation) | 2022 | [33] |
| Yue Xu et al. | Pretrain: 60,000 (virtual library); Fine tuning: 1,200 (in vitro) | mTP | Graph; Descriptors | GIN | Regressor | Pearson correlation coefficients: 0.573 (test set) | 2024 | [34] |
| Bowen Li et al. | 584 (in vitro) | mTP | Descriptors | XGBoost | Classifier | Area under the curve: 0.983 Precision–recall area under the curve: 0.987 (test set) | 2024 | [35] |
| Wei Wang et al. | 351 (in vivo; Literature-curated) | Delivery efficiency Apparent pKa | ECFP | LightGBM | Classifier, Regressor | Delivery efficiency: F1 score 0.78 Apparent pKa: R2 0.59 (test set) | 2024 | [36] |
| Tianhao Yu et al. | Pretrain: 10 million (virtual library)Fine tuning: 1. Experimental data: Not available 2. AGILE: 1,200 (in vitro; Literature-curated) | Multi tasks | SMILES | Transformer Reinforcement learning | Regressor | in-vivo overall fluresence intensity: 0.98 Liver: 0.98 Spleen: 0.95 Lung: 0.92 PDI: 0.97 Encapsulation efficiency: 0.92 Particle size: 0.90 Cr(Toxicity): 0.79 mRNA transfection potency: 0.98 (test set) | 2025 | [37] |
AI: artificial intelligence; PLS: partial least-squares regression; ECFP: extended-connectivity fingerprints; mRNA: messenger RNA; mTP: mRNA transfection potency; SMILES: simplified molecular input line entry system; PDI: polydispersity index; GIN: graph isomorphism network; mTP: mRNA transfection potency; AGILE: AI-guided lipid engineering framework; IgG: immunoglobulin G; LightGBM: light gradient boosting machine; LightGBM: light gradient boosting machine; XGBoost: extreme gradient boosting.
A pioneering study by Dinesh et al. established one of the first predictive models for the apparent pKa of LNPs[32]. By curating a dataset from literature sources[12], where only the headgroups of the lipids were varied, the authors utilized partial least squares (PLS) regression with 10 molecular descriptors to predict pKa. The model achieved an R2 of 0.97 on the training set and a Q2 of 0.83 under cross-validation. Furthermore, when tested on eight newly designed lipids, it maintained a strong correlation of 93.3% between predicted and experimentally measured pKa values. In a separate effort, Wei et al. manually assembled a dataset from literature and patent sources, employing ECFP generated by RDKit and the LightGBM algorithm to build predictive models for both apparent pKa and mRNA delivery efficiency[36]. The pKa prediction model achieved an R2 of 0.59, while the classification model for delivery efficiency attained an F1 score of 0.79. In an earlier related study, the same group also developed a predictor for IgG titers induced by mRNA vaccines using similar features and algorithms, which achieved an R² exceeding 0.87[33].
Moving beyond traditional machine learning, Xu et al. applied a graph isomorphism network (GIN) to screen ionizable lipids with high mRNA transfection capability from a large virtual combinatorial library[34]. Their workflow involved three stages: first, pre-training a GNN model via contrastive learning on a virtual library of 60,000 lipids; second, fine-tuning the model with experimental data from 1,200 synthesized lipids tested in RAW 264.7 and HeLa cells; and third, using an ensemble of five models to evaluate 12,000 candidate lipids. Among the top-performing candidates, lipid H9 demonstrated comparable efficacy to ALC-0315 after intramuscular administration, with improved muscle specificity and a better safety profile (lower alanine aminotransferase/aspartate aminotransferase levels), underscoring the potential of combining deep learning with high-throughput experimental validation. In another study from Bowen’s group, combinatorial chemistry and machine learning were integrated to accelerate ionizable lipid discovery[35]. The team used a four-component reaction to synthesize a training library of 584 lipid samples, with mRNA transfection potency measured in HeLa cells. Molecular descriptors for headgroup, linker, and tail regions were calculated separately and used to train an extreme gradient boosting (XGBoost) model. The resulting predictor showed a receiver operating characteristic of 0.983 and a precision-recall area under the curve of 0.987, enabling the in silico screening of 40,000 virtual lipids. Three of the selected lipids exhibited higher transfection efficiency than benchmark lipids MC3 and SM-102.
Further advancing de novo design, Yu et al. adopted a fragment-based approach in which known ionizable lipids were decomposed into building blocks, and a reinforcement learning method was used to guide reassembly[37]. This process generated a library of 10 million virtual ionizable lipids, which served as the pre-training dataset for LipidBER, a bidirectional encoder representations from transformers (BERT)-based language model tailored for lipid sequences. After pre-training, the model was fine-tuned with wet-lab data to predict key LNP properties such as potency, target specificity, size, polydispersity index (PDI), and safety profile, achieving high predictive accuracy (R2 > 0.9 for most tasks). This method offers a powerful in silico strategy for generating novel ionizable lipids with optimized biological characteristics.
These aforementioned studies cover a wide range of applications of predictive models, from physicochemical properties such as particle size, PDI, apparent pKa, to mRNA transfection efficacy, organ tropism, as well as immunogenicity. Though different studies have employed diverse datasets and methodologies, most have evaluated their models using novel ionizable lipids with desirable potential properties. These findings have bolstered researchers’ confidence in applying AI for the development of structurally novel ionizable lipids with superior performance. However, it should be noted that the generalizability of these models across different administration routes and species hasn’t been validated. In addition, high predictive performance in vitro does not necessarily guarantee in vivo performance, since biological environments are complicated. Therefore, in vivo validation remains indispensable for bridging the gap between computational prediction and actual performance. As more high-quality in vivo data is being accumulated, the precision of these models will continue to improve, enabling researchers to filter desirable ionizable lipids more efficiently. This in silico design and selection of promising compounds not only saves researchers’ time but also conserves experimental resources.
4. Experimental Data Accumulation
LNPs have achieved remarkable clinical success as efficient drug delivery systems. However, effectively targeting non-hepatic organs remains a significant challenge, hampered by biological barriers such as complex tissue microenvironments and the formation of distinct biomolecular coronas in vivo. While most current AI models are trained on in vitro cell data, which is more straightforward to acquire and process, these models often fail to accurately predict LNP behavior in living organisms[38]. This is largely because delivery efficiency differs substantially between controlled in vitro settings and dynamic in vivo environments, where factors like serum protein adsorption and circulatory stability come into play.
In recent years, extensive academic exploration has focused on developing LNPs capable of specifically targeting extrahepatic organs using in vivo screening experiments[39]. Substantial literature confirms that designing novel lipid molecules or adjusting LNP formulation compositions can significantly alter their biodistribution. Key studies include: A 2020 study introduced the concept of selective organ targeting (SORT) lipids, demonstrating that incorporating these lipids to adjust the internal charge of LNPs influences tissue specificity in mice, targeting lungs, spleen, and liver[40]; A 2023 study evaluated 94 LNPs with different PEG-lipids in head and neck squamous cell carcinoma xenograft mouse models, using DNA-barcoded mRNA[41]; A 2022 study by Genetech assessed 54 LNPs with varying PEG-lipids and ratios for targeting cortical neurons in mice[42]; A 2023 study developed a library of 23 ionizable lipids with modified linkers and tails, testing them in mice for liver and macrophage targeting[43]; Xu’s work in 2022 identified N-series LNPs (with amide bonds in the tail) for efficient mRNA delivery to mouse lungs, where modifying the headgroup of these LNPs influenced targeting of different lung cell types[44]. Delivering Tsc2 mRNA using these LNPs reshaped TSC2 tumor suppression in a LAM model, reducing tumor burden[44]. Another study screened 11 lipids to identify 113-O12B for highly specific lymph node targeting in mice[45].
AI-driven discovery of LNPs for novel applications requires substantial amounts of experimental data for model training. Among these, in vivo biodistribution data are especially valuable, as they provide key insights into the mechanisms of action and toxicity of LNPs. The rapid advancement of barcoding and next-generation sequencing (NGS) technologies has empowered researchers to conduct high-throughput screening of LNPs by uniquely tagging them with barcodes, providing a powerful tool for in-depth study of LNP biodistribution in vivo. The evolution of barcoding strategies for LNP screening includes: DNA Barcodes: Early work by Wang et al. used DNA barcodes encapsulated within LNPs[46]. mRNA Barcodes: In 2019, Michael Mitchell at the University of Pennsylvania proposed that mRNA barcodes more accurately represent the physicochemical properties of mRNA-loaded LNPs compared to DNA barcodes[47]. They designed an mRNA barcode concatenated with Luciferase mRNA to study the biodistribution of LNP formulations across eight organs in mice. DNA Barcode + Cre-mRNA: In 2023, James Dahlman’s group at Georgia Tech employed DNA-barcoded LNPs co-encapsulating Cre-mRNA in Ai14 transgenic mice[48]. This allowed high-throughput screening of functional mRNA delivery by detecting tdTomato expression, coupled with single-cell RNA-seq to investigate transcriptomic pathway changes induced by different LNPs at the cellular level. Peptide Barcodes: Also in 2023, Daniel Anderson’s team at MIT developed a peptide barcode technology[49]. This involves encapsulating mRNAs encoding unique peptide sequences into distinct LNPs. Successful delivery and translation can be quantified using liquid chromatography-tandem mass spectrometry. This approach was used to screen 400 LNPs comprising 384 ionizable cationic lipids for functional delivery across multiple organs in mice. Notably, DNA barcodes are physically separated from the cargo mRNAs, a key distinction from mRNA barcodes, which are integrated into the 3’ untranslated region of the cargo mRNAs themselves. Furthermore, peptide barcodes offer a unique advantage in that they can directly report on the translation efficiency of the cargo mRNAs, providing more functional insights into LNP-mediated delivery outcomes.
Integrating barcoding with different sequencing methodologies allows for multi-faceted analysis of LNP biodistribution under various scenarios: RNA-Seq: This high-throughput sequencing technique quantifies the abundance of each barcode, representing the concentration and proportion of each LNP entering specific organs or cells. It helps determine the distribution of LNPs within the body. Polysome-Seq: This technique assesses translational activity by measuring the number of ribosomes bound to each barcode-mRNA, stratifying translating versus non-translating mRNAs. It provides insights into the functional expression of LNP-delivered mRNA during translation. Single-cell RNA-seq (scRNA-seq): This technology enables comprehensive transcriptomic analysis of individual cells. By isolating and sequencing thousands of single cells from a tissue, it annotates cell types based on transcriptomic data. When analyzing barcode content at the single-cell level, scRNA-seq can pinpoint the distribution of LNPs among different cell types, further elucidating their in vivo mechanisms of action. Collectively, these sequencing-based approaches decode distinct biological insights: RNA-Seq reveals the tissue/cell-targeting efficiency of LNPs by quantifying their accumulation specificity; Polysome-Seq directly reflects the functional translation efficacy of delivered mRNA, linking LNP structure to actionable biological output; scRNA-seq uncovers the cell-type-specific distribution and transcriptomic regulatory effects of LNPs at single-cell resolution.
Notably, the large-scale, high-quality in vivo datasets generated by these barcoding-enabled sequencing campaigns lay a solid foundation for AI integration. Machine learning and deep learning algorithms can leverage this multi-dimensional data, encompassing targeting specificity, translation activity, and cell-type selectivity, to build predictive models, such as QSAR models for property prediction, and generative models for novel molecular design, thereby accelerating the rational optimization of ionizable lipids and LNP formulations.
5. Discussion
Although AI shows considerable potential to accelerate the design of ionizable lipids, its practical application in this field still faces several key limitations. First, the predictive performance and generalization capability of AI models are highly dependent on the quality and representativeness of the training data[28]. Biases or inconsistencies in datasets, such as imbalanced data distribution, inaccurately labeled experimental outcomes, or unresolved batch effects, can significantly skew model outputs. Rather than capturing meaningful QSAR, poorly curated data may lead the model to overfit to experimental artifacts or noise, thereby reducing its real-world applicability. Second, although high-throughput screening methods such as barcoding combined with NGS have greatly enhanced the pace of data generation, issues around experimental reproducibility and stability remain pronounced, especially in complex in vivo systems. Variability in animal models, administration protocols, and tissue-processing workflows sets a lower bound on data reliability, which in turn constrains the accuracy and trustworthiness of AI models trained on such data. Third, while established AI algorithms from small-molecule drug discovery can be partially transferred to lipid design, ionizable lipids possess distinctive structural features that necessitate tailored modeling strategies. These molecules typically follow a modular architecture composed of headgroups, linkers, and tails. To ensure generated structures are both synthetically feasible and biologically relevant, AI-based generative models must incorporate domain-specific constraints, such as filtering out unstable substructures and adhering to bio-inspired structural patterns. While this may narrow the theoretical chemical space, it significantly increases the likelihood of identifying practical and effective lipid candidates. In summary, overcoming these barriers, data quality, experimental reproducibility, and structure-aware model design, will be essential for AI to realize its full potential in the rational design of ionizable lipids.
In addition to data-driven AI models, scientific computing approaches provide powerful tools for the rational design of ionizable lipids and the investigation of their mechanisms of action. For instance, molecular docking can be employed to simulate the interactions between lipid molecules and key inflammation-related signaling proteins, thereby aiding in the early assessment of potential safety profiles. Furthermore, molecular dynamics (MD) simulations enable the calculation of critical physicochemical parameters, such as the lipid packing parameter[50], which correlates with endosomal escape efficiency, and can model pH-dependent phase transitions that are crucial for mRNA delivery[51]. These computational insights can be powerfully combined with experimental structural biology techniques. Specifically, small-angle X-ray scattering (SAXS) can be integrated with simulation data to elucidate the internal nanostructure of LNPs[51], providing a deeper understanding of the relationship between lipid composition, self-assembled structure, and biological function. MD evidence indicates that the inverse micellar-to-inverse hexagonal transition of ionizable lipid phases constitutes a rate-limiting step in endosomal release, with the extent of this transition directly correlating with protein expression levels[51]. These approaches deepen understanding of lipid composition, self-assembled structure, and biological function.
In summary, this mini-review highlights how AI and computational approaches are reshaping the discovery and optimization of ionizable lipids by enabling data-driven exploration beyond traditional empirical design. However, the aforementioned limitations still need to be addressed. These early successes suggest that AI has moved beyond proof-of-concept in this field, while also underscoring the importance of realistic expectations regarding model generalizability and translational relevance. Looking forward, continued expansion of high-quality in vivo datasets, advances in structure-aware and physics-informed algorithms, and tighter integration between computation and experimentation are expected to support a more robust and rational paradigm for the development of next-generation ionizable lipids for mRNA therapeutics. Overall, these advances position AI as a promising tool in the rational design of lipid-based delivery systems.
This schematic illustrates an integrated workflow for the discovery and optimization of ionizable lipids using AI and data-driven modeling. (A) Lipid structures are encoded using multiple molecular representations, including physicochemical descriptors, molecular fingerprints, SMILES strings, and graph-based formulations, enabling flexible input formats for different modeling strategies. (B) Representative AI models applied at different stages of lipid design include sequence-based language models for molecular generation (e.g., BERT), reinforcement learning frameworks for goal-directed optimization, and descriptor- or fingerprint-based machine-learning models (e.g., XGBoost) for property prediction and ranking. (C) Large virtual libraries of ionizable lipids are constructed through combinatorial enumeration of modular lipid components and deep learning–based molecular generation approaches. (D) Model training and validation leverage diverse data sources, including curated literature reports, patent-derived datasets, and results from high-throughput lipid screening. (E) Trained predictive models enable in silico screening and prioritization of virtual lipid candidates based on multiple performance-related properties.
Acknowledgements
The authors thank all contributors for helpful discussions. Language polishing was performed with the assistance of an AI-based tool (Doubao). All scientific content was developed by the authors.
Authors contribution
Liang D, Xu C: Conceptualization, methodology, writing-original draft.
Li H, Ma X, Gao P, Ying B: Validation, writing-review & editing.
Conflicts of interest
All authors are employees of Suzhou Abogen Biosciences Inc., Ltd. The authors declare no other competing interests.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Availability of data and materials
Not applicable.
Funding
None.
Copyright
© The Author(s) 2026.
References
-
1. Gote V, Bolla PK, Kommineni N, Butreddy A, Nukala PK, Palakurthi SS, et al. A comprehensive review of mRNA vaccines. Int J Mol Sci. 2023;24(3):2700.[DOI]
-
2. Zhang Y, Sun C, Wang C, Jankovic KE, Dong Y. Lipids and lipid derivatives for RNA delivery. Chem Rev. 2021;121(20):12181-12277.[DOI]
-
3. Cullis PR, Felgner PL. The 60-year evolution of lipid nanoparticles for nucleic acid delivery. Nat Rev Drug Discov. 2024;23(9):709-722.[DOI]
-
4. Eygeris Y, Gupta M, Kim J, Sahay G. Chemistry of lipid nanoparticles for RNA delivery. Acc Chem Res. 2022;55(1):2-12.[DOI]
-
5. Hald Albertsen C, Kulkarni JA, Witzigmann D, Lind M, Petersson K, Simonsen JB. The role of lipid components in lipid nanoparticles for vaccines and gene therapy. Adv Drug Deliv Rev. 2022;188:114416.[DOI]
-
6. Han X, Zhang H, Butowska K, Swingle KL, Alameh MG, Weissman D, et al. An ionizable lipid toolbox for RNA delivery. Nat Commun. 2021;12:7233.[DOI]
-
7. Gyanani V, Goswami R. Key design features of lipid nanoparticles and electrostatic charge-based lipid nanoparticle targeting. Pharmaceutics. 2023;15(4):1184.[DOI]
-
8. Maier MA, Jayaraman M, Matsuda S, Liu J, Barros S, Querbes W, et al. Biodegradable lipids enabling rapidly eliminated lipid nanoparticles for systemic delivery of RNAi therapeutics. Mol Ther. 2013;21(8):1570-1578.[DOI]
-
9. Wasungu L, Hoekstra D. Cationic lipids, lipoplexes and intracellular delivery of genes. J Control Release. 2006;116(2):255-264.[DOI]
-
10. Kulkarni JA, Cullis PR, van der Meel R. Lipid nanoparticles enabling gene therapies: From concepts to clinical utility. Nucleic Acid Ther. 2018;28(3):146-157.[DOI]
-
11. Barbier AJ, Jiang AY, Zhang P, Wooster R, Anderson DG. The clinical progress of mRNA vaccines and immunotherapies. Nat Biotechnol. 2022;40(6):840-854.[DOI]
-
12. Jayaraman M, Ansell SM, Mui BL, Tam YK, Chen J, Du X, et al. Maximizing the potency of siRNA lipid nanoparticles for hepatic gene silencing in vivo. Angew Chem Int Ed. 2012;51(34):8529-8533.[DOI]
-
13. Hashiba K, Taguchi M, Sakamoto S, Otsu A, Maeda Y, Suzuki Y, et al. Impact of lipid tail length on the organ selectivity of mRNA-lipid nanoparticles. Nano Lett. 2024;24(41):12758-12767.[DOI]
-
14. Sabnis S, Kumarasinghe ES, Salerno T, Mihai C, Ketova T, Senn JJ, et al. A novel amino lipid series for mRNA delivery: Improved endosomal escape and sustained pharmacology and safety in non-human Primates. Mol Ther. 2018;26(6):1509-1519.[DOI]
-
15. Ansell SM, Du X, inventors. Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids. World patent WO2017075531 A1. 2017.
-
16. Benenato KE, Kumarasinghe ES, Cornebise M, inventors. Novel lipids and lipid nanoparticle formulations for delivery of nucleic acids. Canadian patent application CA 2998810 A1. 2017.
-
17. Payne JE, Chivukula P, Tanis SP, Karmali P, inventors. Ionizable cationic lipid for RNA delivery. United States patent US 9670152 B2. 2018.
-
18. Han X, Alameh MG, Xu Y, Palanki R, El-Mayta R, Dwivedi G, et al. Optimization of the activity and biodegradability of ionizable lipids for mRNA delivery via directed chemical evolution. Nat Biomed Eng. 2024;8(11):1412-1424.[DOI]
-
19. Luong KD, Singh A. Application of transformers in cheminformatics. J Chem Inf Model. 2024;64(11):4392-4409.[DOI]
-
20. O’Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR. Open Babel: An open chemical toolbox. J Cheminform. 2011;3:33.[DOI]
-
21. Yap CW. PaDEL-descriptor: An open source software to calculate molecular descriptors and fingerprints. J Comput Chem. 2011;32(7):1466-1474.[DOI]
-
22. Rogers D, Hahn M. Extended-connectivity fingerprints. J Chem Inf Model. 2010;50(5):742-754.[DOI]
-
23. Carhart RE, Smith DH, Venkataraghavan R. Atom pairs as molecular features in structure-activity studies: Definition and applications. J Chem Inf Comput Sci. 1985;25(2):64-73.[DOI]
-
24. Durant JL, Leland BA, Henry DR, Nourse JG. Reoptimization of MDL keys for use in drug discovery. J Chem Inf Comput Sci. 2002;42(6):1273-1280.[DOI]
-
25. Weininger D. SMILES, a chemical language and information system. 1. J Chem Inf Comput Sci. 1988;28(1):31-36.[DOI]
-
26. Heller SR, McNaught A, Pletnev I, Stein S, Tchekhovskoi D. InChI, the IUPAC international chemical identifier. J Cheminform. 2015;7:23.[DOI]
-
27. Zhang K, Yang X, Wang Y, Yu Y, Huang N, Li G, et al. Artificial intelligence in drug development. Nat Med. 2025;31(1):45-59.[DOI]
-
28. Vora LK, Gholap AD, Jetha K, Thakur RRS, Solanki HK, Chavda VP. Artificial intelligence in pharmaceutical technology and drug delivery design. Pharmaceutics. 2023;15(7):1916.[DOI]
-
29. Catacutan DB, Alexander J, Arnold A, Stokes JM. Machine learning in preclinical drug discovery. Nat Chem Biol. 2024;20(8):960-973.[DOI]
-
30. Zeng X, Wang F, Luo Y, Kang SG, Tang J, Lightstone FC, et al. Deep generative molecular design reshapes drug discovery. Cell Rep Med. 2022;3(12):100794.[DOI]
-
31. van den Broek RL, Patel S, van Westen GJP, Jespers W, Sherman W. In search of beautiful molecules: A perspective on generative modeling for drug design. J Chem Inf Model. 2025;65(18):9383-9397.[DOI]
-
32. Dhumal DM, Patil PD, Kulkarni RV, Akamanchi KG. Experimentally validated QSAR model for surface pKa prediction of heterolipids having potential as delivery materials for nucleic acid therapeutics. ACS Omega. 2020;5(49):32023-32031.[DOI]
-
33. Wang W, Feng S, Ye Z, Gao H, Lin J, Ouyang D. Prediction of lipid nanoparticles for mRNA vaccines by the machine learning algorithm. Acta Pharm Sin B. 2022;12(6):2950-2962.[DOI]
-
34. Xu Y, Ma S, Cui H, Chen J, Xu S, Gong F, et al. AGILE platform: A deep learning powered approach to accelerate LNP development for mRNA delivery. Nat Commun. 2024;15:6305.[DOI]
-
35. Li B, Raji IO, Gordon AGR, Sun L, Raimondo TM, Oladimeji FA, et al. Accelerating ionizable lipid discovery for mRNA delivery using machine learning and combinatorial chemistry. Nat Mater. 2024;23(7):1002-1008.[DOI]
-
36. Wang W, Chen K, Jiang T, Wu Y, Wu Z, Ying H, et al. Artificial intelligence-driven rational design of ionizable lipids for mRNA delivery. Nat Commun. 2024;15:10804.[DOI]
-
37. Yu T, Yao C, Sun Z, Shi F, Zhang L, Lyu K, et al. LipidBERT: A lipid language model pre-trained on METiS de novo lipid library. arXiv:2408.06150 [Preprint]. 2024.[DOI]
-
38. Yuan Y, Wu Y, Cheng J, Yang K, Xia Y, Wu H, et al. Applications of artificial intelligence to lipid nanoparticle delivery. Particuology. 2024;90:88-97.[DOI]
-
39. Kularatne RN, Crist RM, Stern ST. The future of tissue-targeted lipid nanoparticle-mediated nucleic acid delivery. Pharmaceuticals. 2022;15(7):897.[DOI]
-
40. Cheng Q, Wei T, Farbiak L, Johnson LT, Dilliard SA, Siegwart DJ. Selective organ targeting (SORT) nanoparticles for tissue-specific mRNA delivery and CRISPR–Cas gene editing. Nat Nanotechnol. 2020;15(4):313-320.[DOI]
-
41. Huayamares SG, Lokugamage MP, Rab R, Da Silva Sanchez AJ, Kim H, Radmand A, et al. High-throughput screens identify a lipid nanoparticle that preferentially delivers mRNA to human tumors in vivo. J Control Release. 2023;357:394-403.[DOI]
-
42. Sarode A, Fan Y, Byrnes AE, Hammel M, Hura GL, Fu Y, et al. Predictive high-throughput screening of PEGylated lipids in oligonucleotide-loaded lipid nanoparticles for neuronal gene silencing. Nanoscale Adv. 2022;4(9):2107-2123.[DOI]
-
43. Naidu GS, Yong SB, Ramishetti S, Rampado R, Sharma P, Ezra A, et al. A combinatorial library of lipid nanoparticles for cell type-specific mRNA delivery. Adv Sci. 2023;10(19):2301929.[DOI]
-
44. Qiu M, Tang Y, Chen J, Muriph R, Ye Z, Huang C, et al. Lung-selective mRNA delivery of synthetic lipid nanoparticles for the treatment of pulmonary lymphangioleiomyomatosis. Proc Natl Acad Sci U S A. 2022;119(8):e2116271119.[DOI]
-
45. Chen J, Ye Z, Huang C, Qiu M, Song D, Li Y, et al. Lipid nanoparticle-mediated lymph node–targeting delivery of mRNA cancer vaccine elicits robust CD8+ T cell response. Proc Natl Acad Sci U S A. 2022;119(34):e2207841119.[DOI]
-
46. Dahlman JE, Kauffman KJ, Xing Y, Shaw TE, Mir FF, Dlott CC, et al. Barcoded nanoparticles for high throughput in vivo discovery of targeted therapeutics. Proc Natl Acad Sci U S A. 2017;114(8):2060-2065.[DOI]
-
47. Guimaraes PPG, Zhang R, Spektor R, Tan M, Chung A, Billingsley MM, et al. Ionizable lipid nanoparticles encapsulating barcoded mRNA for accelerated in vivo delivery screening. J Control Release. 2019;316:404-417.[DOI]
-
48. Radmand A, Lokugamage MP, Kim H, Dobrowolski C, Zenhausern R, Loughrey D, et al. The transcriptional response to lung-targeting lipid nanoparticles in vivo. Nano Lett. 2023;23(3):993-1002.[DOI]
-
49. Rhym LH, Manan RS, Koller A, Stephanie G, Anderson DG. Peptide-encoding mRNA barcodes for the high-throughput in vivo screening of libraries of lipid nanoparticles for mRNA delivery. Nat Biomed Eng. 2023;7(7):901-910.[DOI]
-
50. Kobierski J, Wnętrzak A, Chachaj-Brekiesz A, Dynarowicz-Latka P. Predicting the packing parameter for lipids in monolayers with the use of molecular dynamics. Colloids Surf B Biointerfaces. 2022;211:112298.[DOI]
-
51. Philipp J, Dabkowska A, Reiser A, Frank K, Krzysztoń R, Brummer C, et al. pH-dependent structural transitions in cationic ionizable lipid mesophases are critical for lipid nanoparticle function. Proc Natl Acad Sci U S A. 2023;120(50):e2310491120.[DOI]
Copyright
© The Author(s) 2026. This is an Open Access article licensed under a Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, sharing, adaptation, distribution and reproduction in any medium or format, for any purpose, even commercially, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Publisher’s Note
Share And Cite




